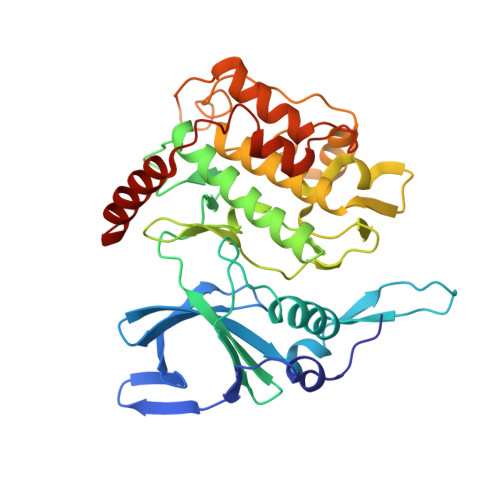
Structures of P. falciparum protein kinase 7 identify an activation motif and leads for inhibitor design.
Merckx, A., Echalier, A., Langford, K., Sicard, A., Langsley, G., Joore, J., Doerig, C., Noble, M., Endicott, J.(2008) Structure 16: 228-238
- PubMed: 18275814
- DOI: https://doi.org/10.1016/j.str.2007.11.014
- Primary Citation Related Structures:
2PMN, 2PMO - PubMed Abstract:
Malaria is a major threat to world health. The identification of parasite targets for drug development is a priority and parasitic protein kinases suggest themselves as suitable targets as many display profound structural and functional divergences from their host counterparts. In this paper, we describe the structure of the orphan protein kinase, Plasmodium falciparum protein kinase 7 (PFPK7). Several Plasmodium protein kinases contain extensive insertions, and the structure of PFPK7 reveals how these may be accommodated as excursions from the canonical eukaryotic protein kinase fold. The constitutively active conformation of PFPK7 is stabilized by a structural motif in which the role of the conserved phosphorylated residue that assists in structuring the activation loop of many protein kinases is played by an arginine residue. We identify two series of PFPK7 ATP-competitive inhibitors and suggest further developments for the design of selective and potent PFPK7 lead compounds as potential antimalarials.
- The Laboratory of Molecular Biophysics, Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, United Kingdom.
Organizational Affiliation: