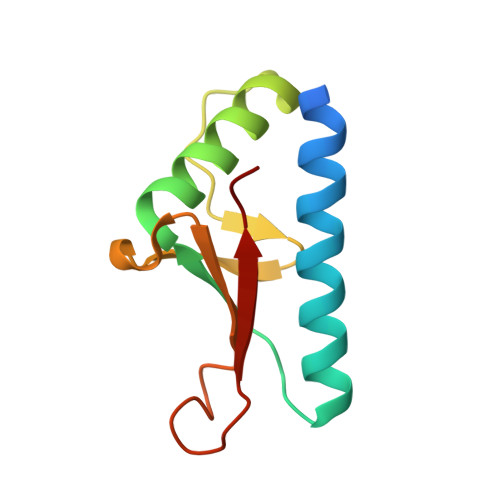
Structure of the hypothetical protein PF0899 from Pyrococcus furiosus at 1.85 A resolution.
Kelley, L.L., Dillard, B.D., Tempel, W., Chen, L., Shaw, N., Lee, D., Newton, M.G., Sugar, F.J., Jenney, F.E., Lee, H.S., Shah, C., Poole, F.L., Adams, M.W., Richardson, J.S., Richardson, D.C., Liu, Z.J., Wang, B.C., Rose, J.(2007) Acta Crystallogr Sect F Struct Biol Cryst Commun 63: 549-552
- PubMed: 17620707 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309107024049
- Primary Citation Related Structures:
2PK8 - PubMed Abstract:
The hypothetical protein PF0899 is a 95-residue peptide from the hyperthermophilic archaeon Pyrococcus furiosus that represents a gene family with six members. P. furiosus ORF PF0899 has been cloned, expressed and crystallized and its structure has been determined by the Southeast Collaboratory for Structural Genomics (http://www.secsg.org). The structure was solved using the SCA2Structure pipeline from multiple data sets and has been refined to 1.85 A against the highest resolution data set collected (a presumed gold derivative), with a crystallographic R factor of 21.0% and R(free) of 24.0%. The refined structure shows some structural similarity to a wedge-shaped domain observed in the structure of the major capsid protein from bacteriophage HK97, suggesting that PF0899 may be a structural protein.
- Southeast Collaboratory for Structural Genomics, Department of Biochemistry and Molecular Biology, University of Georgia, Athens, Georgia, USA.
Organizational Affiliation: