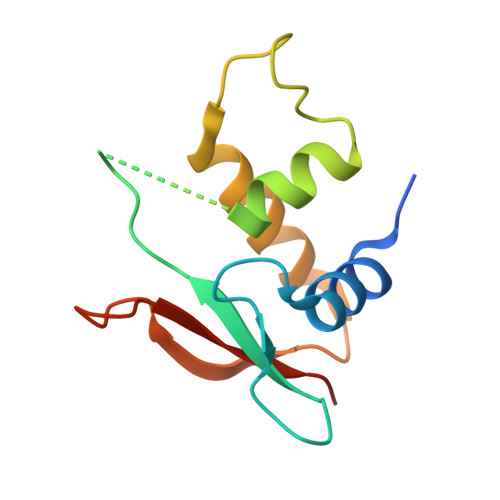
Structure of IRF-3 bound to the PRDIII-I regulatory element of the human interferon-beta enhancer.
Escalante, C.R., Nistal-Villan, E., Shen, L., Garcia-Sastre, A., Aggarwal, A.K.(2007) Mol Cell 26: 703-716
- PubMed: 17560375 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2007.04.022
- Primary Citation Related Structures:
2PI0 - PubMed Abstract:
Interferon regulatory factor 3 (IRF-3) is a key transcription factor in the assembly of the mammalian interferon-beta (IFN-beta) enhanceosome. We present here the structure of IRF-3 DNA binding domain in complex with the complete PRDIII-I regulatory element of the human IFN-beta enhancer. We show that four IRF-3 molecules bind in tandem to, variably spaced, consensus and nonconsensus IRF sites on the composite element. The ability of IRF-3 to bind these variable sites derives in part from two nonconserved arginines (Arg78 and Arg86) that partake in alternate protein-DNA contacts. We also show that the protein-DNA contacts are highly overlapped and that all four IRF sites are required for gene activation in vivo. In addition, we show that changing the nonconsensus IRF sites to consensus sites creates a more efficient enhancer in vivo. Together, the structure and accompanying biological data provide insights into the assembly of the IFN-beta enhanceosome in mammals.
- Department of Structural and Chemical Biology, Mount Sinai School of Medicine, New York, NY 10029, USA.
Organizational Affiliation: