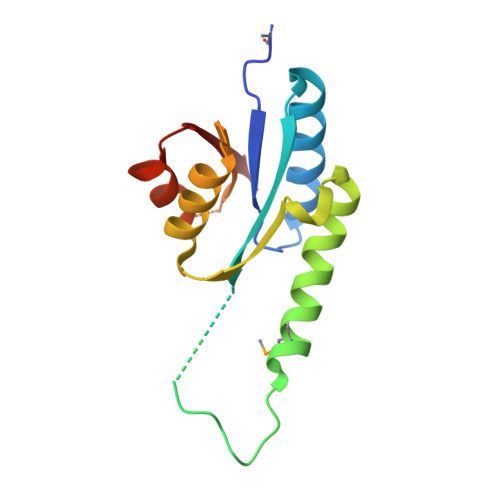
Structural and functional insight into the universal stress protein family.
Tkaczuk, K.L., Shumilin, I.A., Chruszcz, M., Evdokimova, E., Savchenko, A., Minor, W.(2013) Evol Appl 6: 434-449
- PubMed: 23745136 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/eva.12057
- Primary Citation Related Structures:
2PFS, 3QTB, 3TNJ, 6HCD - PubMed Abstract:
We present the crystal structures of two universal stress proteins (USP) from Archaeoglobus fulgidus and Nitrosomonas europaea in both apo- and ligand-bound forms. This work is the first complete synthesis of the structural properties of 26 USP available in the Protein Data Bank, over 75% of which were determined by structure genomics centers with no additional information provided. The results of bioinformatic analyses of all available USP structures and their sequence homologs revealed that these two new USP structures share overall structural similarity with structures of USPs previously determined. Clustering and cladogram analyses, however, show how they diverge from other members of the USP superfamily and show greater similarity to USPs from organisms inhabiting extreme environments. We compared them with other archaeal and bacterial USPs and discuss their similarities and differences in context of structure, sequential motifs, and potential function. We also attempted to group all analyzed USPs into families, so that assignment of the potential function to those with no experimental data available would be possible by extrapolation.
- Department of Molecular Physiology and Biological Physics, University of Virginia Charlottesville, VA, USA ; Midwest Center for Structural Genomics USA.
Organizational Affiliation: