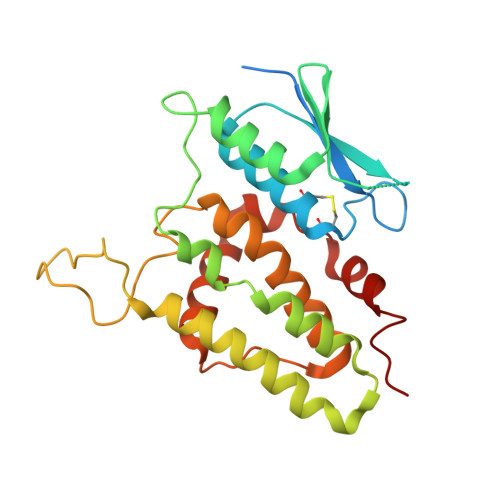
The crystal structure of human chloride intracellular channel protein 2: A disulfide bond with functional implications.
Mi, W., Liang, Y.H., Li, L., Su, X.D.(2008) Proteins 71: 509-513
- PubMed: 18186468 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21922
- Primary Citation Related Structures:
2PER - National Laboratory of Protein Engineering and Plant Genetic Engineering, College of Life Sciences, Peking University, Beijing 100871, China.
Organizational Affiliation: