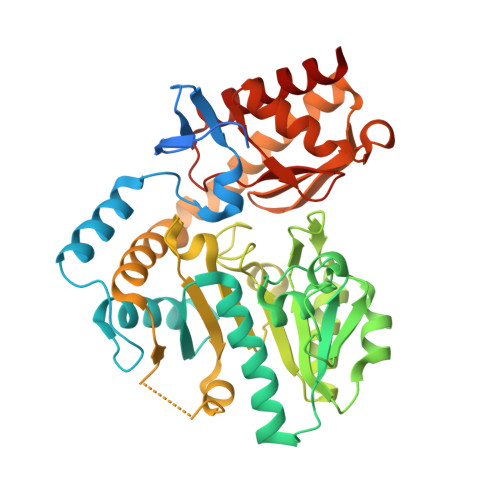
Structure of biosynthetic N-acetylornithine aminotransferase from Salmonella typhimurium: Studies on substrate specificity and inhibitor binding
Rajaram, V., Ratna Prasuna, P., Savithri, H.S., Murthy, M.R.N.(2007) Proteins 70: 429-441
- PubMed: 17680699 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21567
- Primary Citation Related Structures:
2PB0, 2PB2 - PubMed Abstract:
Acetylornithine aminotransferase (AcOAT) is one of the key enzymes involved in arginine metabolism and catalyzes the conversion of N-acetylglutamate semialdehyde to N-acetylornithine (AcOrn) in the presence of L-glutamate. It belongs to the Type I subgroup II family of pyridoxal 5'-phosphate (PLP) dependent enzymes. E. coli biosynthetic AcOAT (eAcOAT) also catalyzes the conversion of N-succinyl-L-2-amino-6-oxopimelate to N-succinyl-L,L-diaminopimelate, one of the steps in lysine biosynthesis. In view of the critical role of AcOAT in lysine and arginine biosynthesis, structural studies were initiated on the enzyme from S. typhimurium (sAcOAT). The K(m) and k(cat)/K(m) values determined with the purified sAcOAT suggested that the enzyme had much higher affinity for AcOrn than for ornithine (Orn) and was more efficient than eAcOAT. sAcOAT was inhibited by gabaculine (Gcn) with an inhibition constant (K(i)) of 7 microM and a second-order rate constant (k(2)) of 0.16 mM(-1) s(-1). sAcOAT, crystallized in the unliganded form and in the presence of Gcn or L-glutamate, diffracted to a maximum resolution of 1.90 A and contained a dimer in the asymmetric unit. The structure of unliganded sAcOAT showed significant electron density for PLP in only one of the subunits (subunit A). The asymmetry in PLP binding could be attributed to the ordering of the loop L(alphak-) (betam) in only one subunit (subunit B; the loop from subunit B comes close to the phosphate group of PLP in subunit A). Structural and spectral studies of sAcOAT with Gcn suggested that the enzyme might have a low affinity for PLP-Gcn complex. Comparison of sAcOAT with T. thermophilus AcOAT and human ornithine aminotransferase suggested that the higher specificity of sAcOAT towards AcOrn may not be due to specific changes in the active site residues but could result from minor conformational changes in some of them. This is the first structural report of AcOAT from a mesophilic organism and could serve as a basis for drug design as the enzyme is important for bacterial cell wall biosynthesis.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560012, India.
Organizational Affiliation: