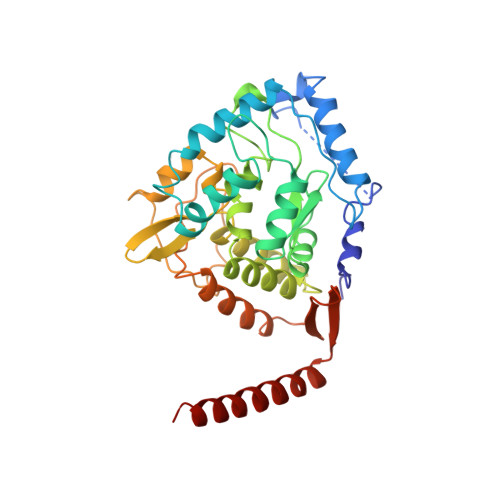
Structure of tetrameric human phenylalanine hydroxylase and its implications for phenylketonuria.
Fusetti, F., Erlandsen, H., Flatmark, T., Stevens, R.C.(1998) J Biological Chem 273: 16962-16967
- PubMed: 9642259 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.273.27.16962
- Primary Citation Related Structures:
2PAH - PubMed Abstract:
Phenylalanine hydroxylase (PheOH) catalyzes the conversion of L-phenylalanine to L-tyrosine, the rate-limiting step in the oxidative degradation of phenylalanine. Mutations in the human PheOH gene cause phenylketonuria, a common autosomal recessive metabolic disorder that in untreated patients often results in varying degrees of mental retardation. We have determined the crystal structure of human PheOH (residues 118-452). The enzyme crystallizes as a tetramer with each monomer consisting of a catalytic and a tetramerization domain. The tetramerization domain is characterized by the presence of a domain swapping arm that interacts with the other monomers forming an antiparallel coiled-coil. The structure is the first report of a tetrameric PheOH and displays an overall architecture similar to that of the functionally related tyrosine hydroxylase. In contrast to the tyrosine hydroxylase tetramer structure, a very pronounced asymmetry is observed in the phenylalanine hydroxylase, caused by the occurrence of two alternate conformations in the hinge region that leads to the coiled-coil helix. Examination of the mutations causing PKU shows that some of the most frequent mutations are located at the interface of the catalytic and tetramerization domains. Their effects on the structural and cellular stability of the enzyme are discussed.
- Department of Chemistry, University of California and Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, California 94720, USA.
Organizational Affiliation: