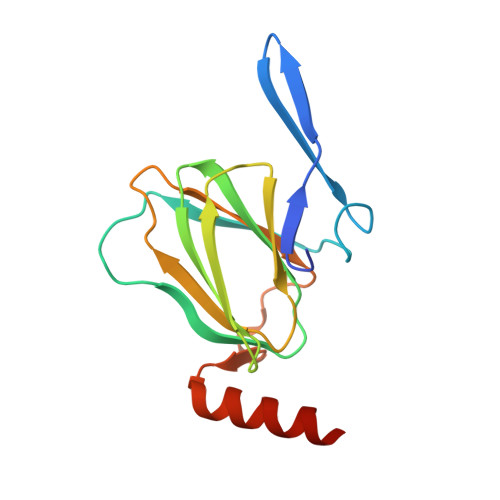
The X-ray Structure of dTDP-4-Keto-6-deoxy-D-glucose-3,4-ketoisomerase.
Davis, M.L., Thoden, J.B., Holden, H.M.(2007) J Biological Chem 282: 19227-19236
- PubMed: 17459872 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M702529200
- Primary Citation Related Structures:
2PA7, 2PAE, 2PAK, 2PAM - PubMed Abstract:
The repeating unit of the glycan chain in the S-layer of the bacterium Aneurinibacillus thermoaerophilus L420-91(T) is composed of four alpha-d-rhamnose molecules and two 3-acetamido-3,6-dideoxy-alpha-d-galactose moieties (abbreviated as Fucp3NAc). Formation of the glycan layer requires nucleotide-activated sugars as the donor molecules. Whereas the enzymes involved in the synthesis of GDP-rhamnose have been well characterized, less is known regarding the structures and enzymatic mechanisms of the enzymes required for the production of dTDP-Fucp3NAc. One of the enzymes involved in the biosynthesis of dTDP-Fucp3NAc is a 3,4-ketoisomerase, hereafter referred to as FdtA. Here we describe the first three-dimensional structure of this sugar isomerase complexed with dTDP and solved to 1.5 A resolution. The FdtA dimer assumes an almost jellyfish-like appearance with the sole alpha-helices representing the tentacles. Formation of the FdtA dimer represents a classical example of domain swapping whereby beta-strands 2 and 3 from one subunit form part of a beta-sheet in the second subunit. The active site architecture of FdtA is characterized by a cluster of three histidine residues, two of which, His(49) and His(51), appear to be strictly conserved in the amino acid sequences deposited to date. Site-directed mutagenesis experiments, enzymatic assays, and x-ray crystallographic analyses suggest that His(49) functions as an active site base.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706-1544, USA.
Organizational Affiliation: