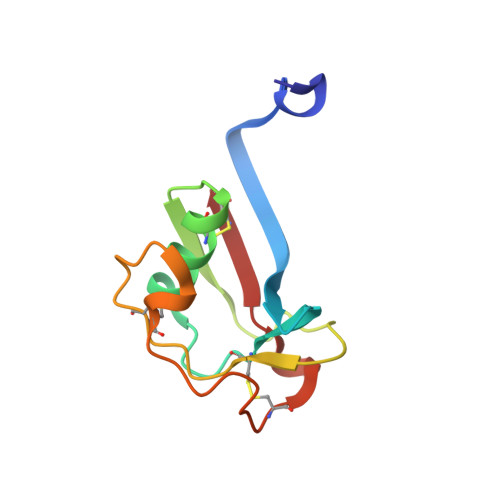
Crystal structure of the cysteine-rich domain of scavenger receptor MARCO reveals the presence of a basic and an acidic cluster that both contribute to ligand recognition.
Ojala, J.R., Pikkarainen, T., Tuuttila, A., Sandalova, T., Tryggvason, K.(2007) J Biological Chem 282: 16654-16666
- PubMed: 17405873 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M701750200
- Primary Citation Related Structures:
2OY3, 2OYA - PubMed Abstract:
MARCO is a trimeric class A scavenger receptor of macrophages and dendritic cells that recognizes polyanionic particles and pathogens. The distal, scavenger receptor cysteine-rich (SRCR) domain of the extracellular part of this receptor has been implicated in ligand binding. To provide a structural basis for understanding the ligand-binding mechanisms of MARCO, we have determined the crystal structure of the mouse MARCO SRCR domain. The recombinant SRCR domain purified as monomeric and dimeric forms, and their structures were determined at 1.78 and 1.77 A resolution, respectively. The monomer has a compact globular fold with a twisted five-stranded antiparallel beta-sheet and a long loop covering a single alpha-helix, whereas the dimer is formed via beta-strand swapping of two monomers, thus containing a large eight-stranded beta-sheet. Calculation of the surface electrostatic potential revealed that the beta-sheet region with several arginines forms a basic cluster. Unexpectedly, an acidic cluster was found in the long loop region. In the monomer, the acidic cluster is involved in metal ion binding. Studies with cells expressing various SRCR domain mutants showed that all of the arginines of the basic cluster are involved in ligand binding, suggesting a cooperative binding mechanism. Ligand binding is also dependent on the acidic cluster and Ca2+ ions whose depletion appears to affect ligand binding at least by modulating the electrostatic potential or relative domain orientation. We propose that the SRCR domain dimerization can contribute to the recognition of large ligands by providing a means for the MARCO receptor oligomerization.
- Division of Matrix Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-17177 Stockholm, Sweden.
Organizational Affiliation: