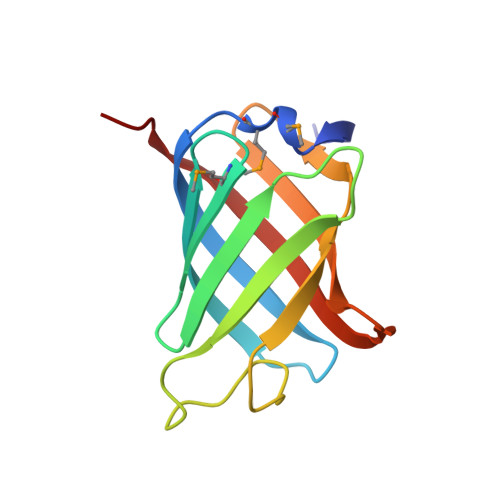
Structure of GrlR and the implication of its EDED motif in mediating the regulation of type III secretion system in EHEC.
Jobichen, C., Li, M., Yerushalmi, G., Tan, Y.W., Mok, Y.K., Rosenshine, I., Leung, K.Y., Sivaraman, J.(2007) PLoS Pathog 3: e69-e69
- PubMed: 17511515
- DOI: https://doi.org/10.1371/journal.ppat.0030069
- Primary Citation of Related Structures:
2OVS - PubMed Abstract:
Enterohemorrhagic Escherichia coli (EHEC) is a common cause of severe hemorrhagic colitis. EHEC's virulence is dependent upon a type III secretion system (TTSS) encoded by 41 genes. These genes are organized in several operons clustered in the locus of enterocyte effacement. Most of the locus of enterocyte effacement genes, including grlA and grlR, are positively regulated by Ler, and Ler expression is positively and negatively modulated by GrlA and GrlR, respectively. However, the molecular basis for the GrlA and GrlR activity is still elusive. We have determined the crystal structure of GrlR at 1.9 A resolution. It consists of a typical beta-barrel fold with eight beta-strands containing an internal hydrophobic cavity and a plug-like loop on one side of the barrel. Strong hydrophobic interactions between the two beta-barrels maintain the dimeric architecture of GrlR. Furthermore, a unique surface-exposed EDED (Glu-Asp-Glu-Asp) motif is identified to be critical for GrlA-GrlR interaction and for the repressive activity of GrlR. This study contributes a novel molecular insight into the mechanism of GrlR function.
- Department of Biological Sciences, National University of Singapore, Singapore.
Organizational Affiliation: