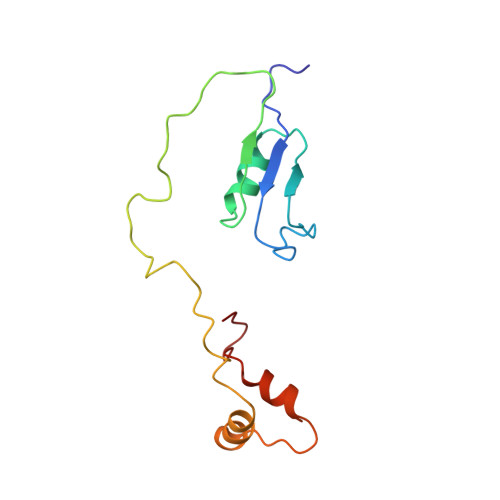
Solution structure of HI1506, a novel two-domain protein from Haemophilus influenzae.
Sari, N., He, Y., Doseeva, V., Surabian, K., Ramprakash, J., Schwarz, F., Herzberg, O., Orban, J.(2007) Protein Sci 16: 977-982
- PubMed: 17400915 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.072820907
- Primary Citation Related Structures:
2OUT - PubMed Abstract:
HI1506 is a 128-residue hypothetical protein of unknown function from Haemophilus influenzae. It was originally annotated as a shorter 85-residue protein, but a more detailed sequence analysis conducted in our laboratory revealed that the full-length protein has an additional 43 residues on the C terminus, corresponding with a region initially ascribed to HI1507. As part of a larger effort to understand the functions of hypothetical proteins from Gram-negative bacteria, and H. influenzae in particular, we report here the three-dimensional solution NMR structure for the corrected full-length HI1506 protein. The structure consists of two well-defined domains, an alpha/beta 50-residue N-domain and a 3-alpha 32-residue C-domain, separated by an unstructured 30-residue linker. Both domains have positively charged surface patches and weak structural homology with folds that are associated with RNA binding, suggesting a possible functional role in binding distal nucleic acid sites.
- Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, Rockville, Maryland 20850, USA.
Organizational Affiliation: