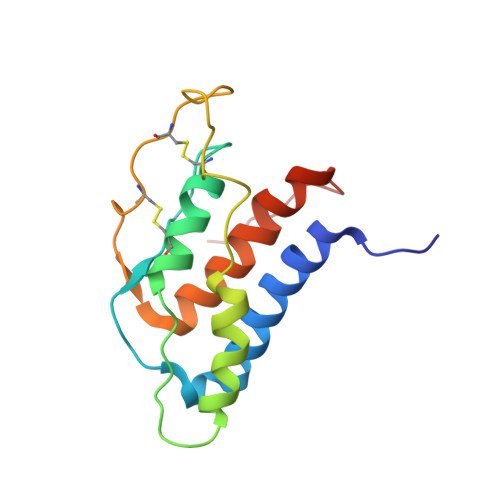
The existence of multiple conformers of interleukin-21 directs engineering of a superpotent analogue.
Bondensgaard, K., Breinholt, J., Madsen, D., Omkvist, D.H., Kang, L., Worsaae, A., Becker, P., Schiodt, C.B., Hjorth, S.A.(2007) J Biological Chem 282: 23326-23336
- PubMed: 17565991 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M701313200
- Primary Citation Related Structures:
2OQP - PubMed Abstract:
The high resolution three-dimensional structure of human interleukin (hIL)-21 has been resolved by heteronuclear NMR spectroscopy. Overall, the hIL-21 structure is dominated by a well defined central four-helical bundle, arranged in an up-up-down-down topology, as observed for other cytokines. A segment of the hIL-21 molecule that includes the third helical segment, helix C, is observed to exist in two distinct and interchangeable states. In one conformer, the helix C segment is presented in a regular, alpha-helical conformation, whereas in the other conformer, this segment is largely disordered. A structure-based sequence alignment of hIL-21 with receptor complexes of the related cytokines, interleukin-2 and -4, implied that this particular segment is involved in receptor binding. An hIL-21 analog was designed to stabilize the region around helix C through the introduction of a segment grafted from hIL-4. This novel hIL-21 analog was demonstrated to exhibit a 10-fold increase in potency in a cellular assay.
- Novo Nordisk A/S, Biopharmaceuticals Research Unit, DK-2760 Måløv, Denmark. kbaa@novonordisk.com
Organizational Affiliation: