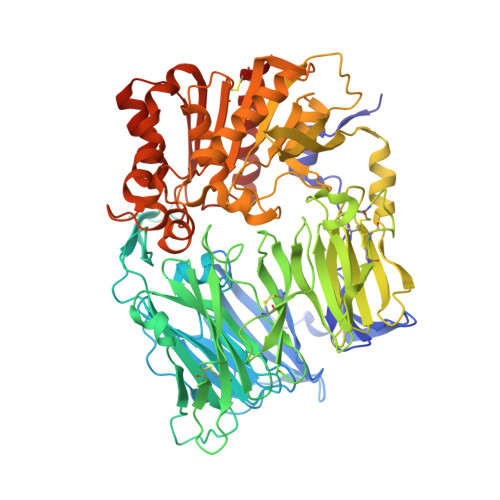
Discovery and Structure-Activity Relationships of Piperidinone- and Piperidine-Constrained Phenethylamines as Novel, Potent, and Selective Dipeptidyl Peptidase IV Inhibitors.
Pei, Z., Li, X., Geldern, T.W., Longenecker, K., Pireh, D., Stewart, K.D., Backes, B.J., Lai, C., Lubben, T.H., Ballaron, S.J., Beno, D.W., Kempf-Grote, A.J., Sham, H.L., Trevillyan, J.M.(2007) J Med Chem 50: 1983-1987
- PubMed: 17367123 Search on PubMed
- DOI: https://doi.org/10.1021/jm061436d
- Primary Citation Related Structures:
2OQI, 2OQV - PubMed Abstract:
Dipeptidyl peptidase IV (DPP4) inhibitors are emerging as a new class of therapeutic agents for the treatment of type 2 diabetes. They exert their beneficial effects by increasing the levels of active glucagon-like peptide-1 and glucose-dependent insulinotropic peptide, which are two important incretins for glucose homeostasis. Starting from a high-throughput screening hit, we were able to identify a series of piperidinone- and piperidine-constrained phenethylamines as novel DPP4 inhibitors. Optimized compounds are potent, selective, and have good pharmacokinetic profiles.
- Metabolic Disease Research, Global Pharmaceutical Research and Development, Abbott Laboratories, 100 Abbott Park Road, Abbott Park, IL 60064-3500, USA. zhonghua.pei@abbott.com
Organizational Affiliation: