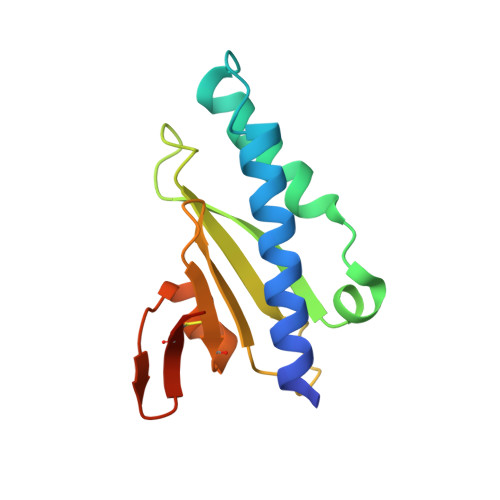
3D structure/function analysis of PilX reveals how minor pilins can modulate the virulence properties of type IV pili.
Helaine, S., Dyer, D.H., Nassif, X., Pelicic, V., Forest, K.T.(2007) Proc Natl Acad Sci U S A 104: 15888-15893
- PubMed: 17893339 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0707581104
- Primary Citation Related Structures:
2OPD, 2OPE - PubMed Abstract:
Type IV pili (Tfp) are widespread filamentous bacterial organelles that mediate multiple virulence-related phenotypes. They are composed mainly of pilin subunits, which are processed before filament assembly by dedicated prepilin peptidases. Other proteins processed by these peptidases, whose molecular nature and mode of action remain enigmatic, play critical roles in Tfp biology. We have performed a detailed structure/function analysis of one such protein, PilX from Neisseria meningitidis, which is crucial for formation of bacterial aggregates and adhesion to human cells. The x-ray crystal structure of PilX reveals the alpha/beta roll fold shared by all pilins, and we show that this protein colocalizes with Tfp. These observations suggest that PilX is a minor, or low abundance, pilin that assembles within the filaments in a similar way to pilin. Deletion of a PilX distinctive structural element, which is predicted to be exposed on the filament surface, abolishes aggregation and adhesion. Our results support a model in which surface-exposed motifs in PilX subunits stabilize bacterial aggregates against the disruptive force of pilus retraction and illustrate how a minor pilus component can enhance the functional properties of pili of rather simple composition and structure.
- Institut National de la Santé et de la Recherche Médicale, U570, 75015 Paris, France.
Organizational Affiliation: