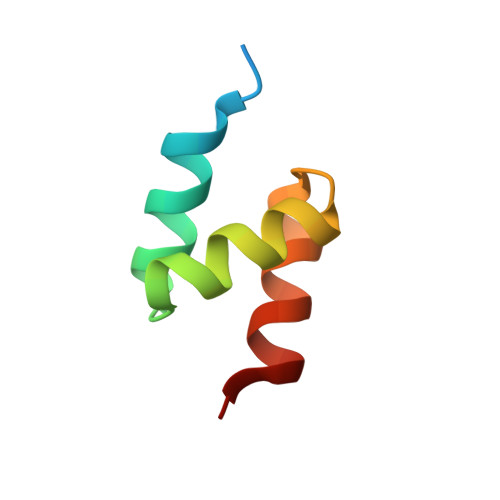
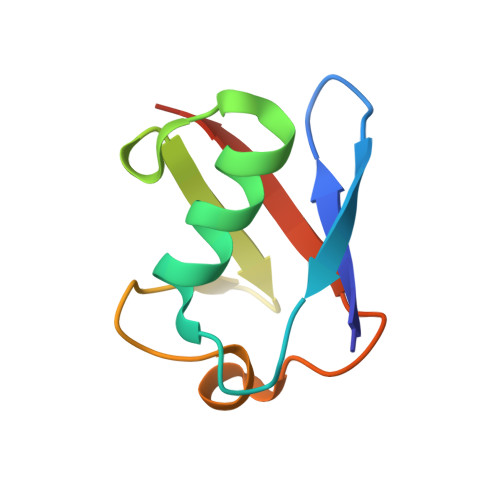
Structural basis for ubiquitin-mediated dimerization and activation of the ubiquitin protein ligase Cbl-b.
Peschard, P., Kozlov, G., Lin, T., Mirza, I.A., Berghuis, A.M., Lipkowitz, S., Park, M., Gehring, K.(2007) Mol Cell 27: 474-485
- PubMed: 17679095 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2007.06.023
- Primary Citation Related Structures:
2OOA, 2OOB - PubMed Abstract:
Cbl proteins are E3 ubiquitin ligases that are negative regulators of many receptor tyrosine kinases. Cbl-b and c-Cbl contain a ubiquitin-associated (UBA) domain, which is present in a variety of proteins involved in ubiquitin-mediated processes. Despite high sequence identity, Cbl UBA domains display remarkably different ubiquitin-binding properties. Here, we report the crystal structure of the UBA domain of Cbl-b in complex with ubiquitin at 1.9 A resolution. The structure reveals an atypical mechanism of ubiquitin recognition by the first helix of the UBA. Helices 2 and 3 of the UBA domain form a second binding surface, which mediates UBA dimerization in the crystal and in solution. Site-directed mutagenesis demonstrates that Cbl-b dimerization is regulated by ubiquitin binding and required for tyrosine phosphorylation of Cbl-b and ubiquitination of Cbl-b substrates. These studies demonstrate a role for ubiquitin in regulating biological activity by promoting protein dimerization.
- Department of Biochemistry, McGill University, Montréal, Canada.
Organizational Affiliation: