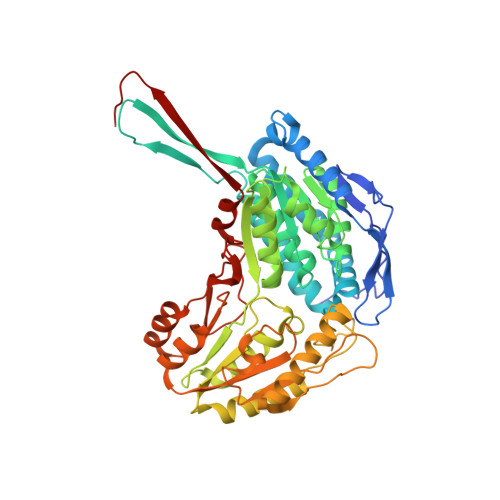
Structural and functional consequences of coenzyme binding to the inactive asian variant of mitochondrial aldehyde dehydrogenase: roles of residues 475 and 487.
Larson, H.N., Zhou, J., Chen, Z., Stamler, J.S., Weiner, H., Hurley, T.D.(2007) J Biological Chem 282: 12940-12950
- PubMed: 17327228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M607959200
- Primary Citation Related Structures:
2ONM, 2ONN, 2ONO, 2ONP - PubMed Abstract:
The common mitochondrial aldehyde dehydrogenase (ALDH2) ALDH2(*)2 polymorphism is associated with impaired ethanol metabolism and decreased efficacy of nitroglycerin treatment. These physiological effects are due to the substitution of Lys for Glu-487 that reduces the k(cat) for these processes and increases the K(m) for NAD(+), as compared with ALDH2. In this study, we sought to understand the nature of the interactions that give rise to the loss of structural integrity and low activity in ALDH2(*)2 even when complexed with coenzyme. Consequently, we have solved the crystal structure of ALDH2(*)2 complexed with coenzyme to 2.5A(.) We have also solved the structures of a mutated form of ALDH2 where Arg-475 is replaced by Gln (R475Q). The structural and functional properties of the R475Q enzyme are intermediate between those of wild-type and the ALDH2(*)2 enzymes. In both cases, the binding of coenzyme restores most of the structural deficits observed in the apoenzyme structures. The binding of coenzyme to the R475Q enzyme restores its structure and catalytic properties to near wild-type levels. In contrast, the disordered helix within the coenzyme binding pocket of ALDH2(*)2 is reordered, but the active site is only partially reordered. Consistent with the structural data, ALDH2(*)2 showed a concentration-dependent increase in esterase activity and nitroglycerin reductase activity upon addition of coenzyme, but the levels of activity do not approach those of the wild-type enzyme or that of the R475Q enzyme. The data presented shows that Glu-487 maintains a critical function in linking the structure of the coenzyme-binding site to that of the active site through its interactions with Arg-264 and Arg-475, and in doing so, creates the stable structural scaffold conducive to catalysis.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, Indiana 46202, USA.
Organizational Affiliation: