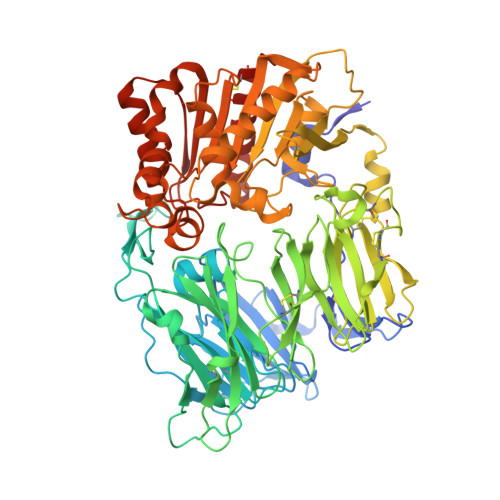
Discovery of alogliptin: a potent, selective, bioavailable, and efficacious inhibitor of dipeptidyl peptidase IV.
Feng, J., Zhang, Z., Wallace, M.B., Stafford, J.A., Kaldor, S.W., Kassel, D.B., Navre, M., Shi, L., Skene, R.J., Asakawa, T., Takeuchi, K., Xu, R., Webb, D.R., Gwaltney, S.L.(2007) J Med Chem 50: 2297-2300
- PubMed: 17441705 Search on PubMed
- DOI: https://doi.org/10.1021/jm070104l
- Primary Citation Related Structures:
2ONC - PubMed Abstract:
Alogliptin is a potent, selective inhibitor of the serine protease dipeptidyl peptidase IV (DPP-4). Herein, we describe the structure-based design and optimization of alogliptin and related quinazolinone-based DPP-4 inhibitors. Following an oral dose, these noncovalent inhibitors provide sustained reduction of plasma DPP-4 activity and a lowering of blood glucose in animal models of diabetes. Alogliptin is currently undergoing phase III trials in patients with type 2 diabetes.