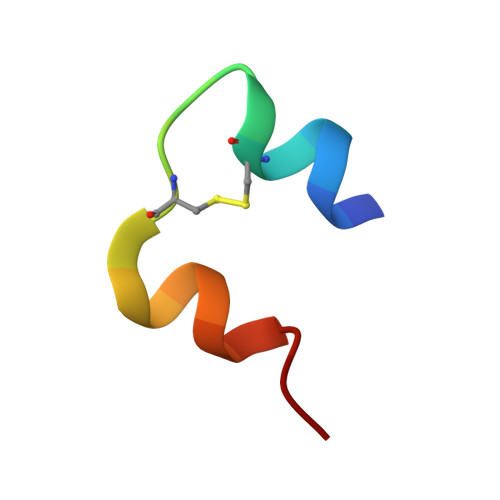
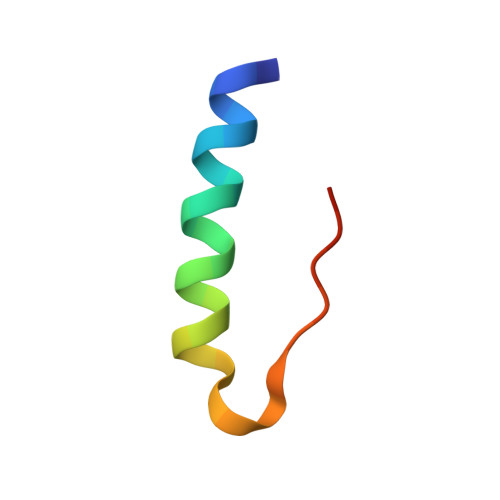
Structural characterization of insulin NPH formulations.
Norrman, M., Hubalek, F., Schluckebier, G.(2007) Eur J Pharm Sci 30: 414-423
- PubMed: 17339105 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejps.2007.01.003
- Primary Citation Related Structures:
2OMG, 2OMH, 2OMI - PubMed Abstract:
Insulin NPH (neutral protamine hagedorn) has for long been one of the most important therapeutic formulations for the treatment of diabetes. The protracted action profile of NPH formulations is gained from crystallizing insulin with zinc in the presence of the basic poly-arginine peptide protamine. In spite of its long history and successful use, the binding mode of the insulin-protamine complex is not known. In this study, three different systems were used to study protamine binding to insulin. In the first system, crystals of an insulin-protamine complex grown in the presence of urea and diffracting to 1.5A resolution were analyzed. In the second system, a shorter peptide consisting of 12 arginine residues was co-crystallized with insulin in order to reduce the flexibility and thereby improve the electron density of the peptide. Both systems yielded data to a significantly higher resolution than obtained previously. In addition, a third system was analyzed where crystals of insulin and protamine were grown in the absence of urea, with conditions closely resembling the pharmaceutical formulation. Data from these NPH microcrystals could for the first time be collected to 2.2A resolution at a micro focused X-ray beamline. Analysis of all three crystal forms reveal potential protamine density located close to the solvent channel leading to the centrally located zinc atoms in the insulin hexamer and support that protamine binds to insulin in a not well defined conformation.
- Diabetes Protein Engineering, Novo Nordisk A/S, Novo Nordisk Park, 2760 Måløv, Denmark.
Organizational Affiliation: