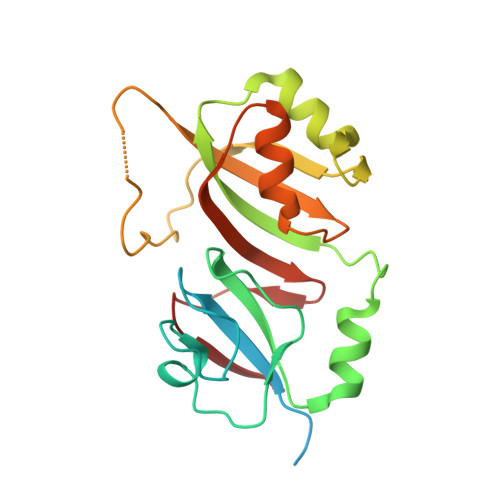
The Crystal Structure of E. coli rRNA Pseudouridine Synthase RluE.
Pan, H., Ho, J.D., Stroud, R.M., Finer-Moore, J.(2007) J Mol Biology 367: 1459-1470
- PubMed: 17320904 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.01.084
- Primary Citation Related Structures:
2OLW, 2OML - PubMed Abstract:
Pseudouridine synthase RluE modifies U2457 in a stem of 23 S RNA in Escherichia coli. This modification is located in the peptidyl transferase center of the ribosome. We determined the crystal structures of the C-terminal, catalytic domain of E. coli RluE at 1.2 A resolution and of full-length RluE at 1.6 A resolution. The crystals of the full-length enzyme contain two molecules in the asymmetric unit and in both molecules the N-terminal domain is disordered. The protein has an active site cleft, conserved in all other pseudouridine synthases, that contains invariant Asp and Tyr residues implicated in catalysis. An electropositive surface patch that covers the active site cleft is just wide enough to accommodate an RNA stem. The RNA substrate stem can be docked to this surface such that the catalytic Asp is adjacent to the target base, and a conserved Arg is positioned to help flip the target base out of the stem into the enzyme active site. A flexible RluE specific loop lies close to the conserved region of the stem in the model, and may contribute to substrate specificity. The stem alone is not a good RluE substrate, suggesting RluE makes additional interactions with other regions in the ribosome.
- Department of Biochemistry and Biophysics, University of California at San Francisco, San Francisco, CA 94143, USA.
Organizational Affiliation: