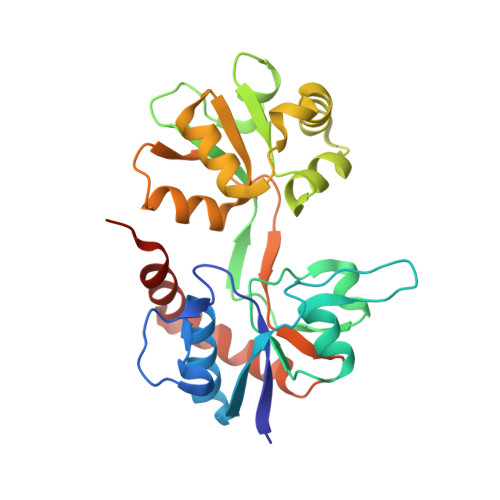
Structure and mechanism of kainate receptor modulation by anions.
Plested, A.J., Mayer, M.L.(2007) Neuron 53: 829-841
- PubMed: 17359918 Search on PubMed
- DOI: https://doi.org/10.1016/j.neuron.2007.02.025
- Primary Citation Related Structures:
2OJT - PubMed Abstract:
L-glutamate, the major excitatory neurotransmitter in the human brain, activates a family of ligand-gated ion channels, the major subtypes of which are named AMPA, kainate, and NMDA receptors. In common with many signal transduction proteins, glutamate receptors are modulated by ions and small molecules, including Ca(2+), Mg(2+), Zn(2+), protons, polyamines, and steroids. Strikingly, the activation of kainate receptors by glutamate requires the presence of both Na(+) and Cl(-) in the extracellular solution, and in the absence of these ions, receptor activity is abolished. Here, we identify the site and mechanism of action of anions. Surprisingly, we find that Cl(-) ions are essential structural components of kainate receptors. Cl(-) ions bind in a cavity formed at the interface between subunits in a dimer pair. In the absence of Cl(-), dimer stability is reduced, the rate of desensitization increases, and the fraction of receptors competent for activation by glutamate drops precipitously.
- Laboratory of Cellular and Molecular Neurophysiology, Porter Neuroscience Research Center, National Institute of Child Health and Human Development, National Institutes of Health, Department of Health and Human Services, Bethesda, MD 20892, USA.
Organizational Affiliation: