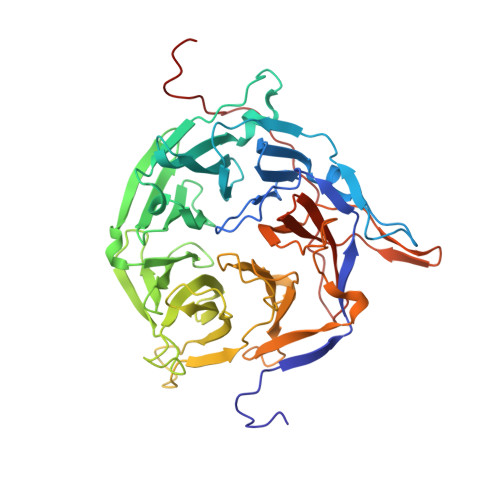
Crystal structure of the N-terminal domain of the human protooncogene Nup214/CAN.
Napetschnig, J., Blobel, G., Hoelz, A.(2007) Proc Natl Acad Sci U S A 104: 1783-1788
- PubMed: 17264208 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0610828104
- Primary Citation Related Structures:
2OIT - PubMed Abstract:
The mammalian nuclear pore complex (NPC) is an approximately 120-MDa proteinaceous assembly consisting of approximately 30 proteins and is the sole gate in the nuclear envelope. The human protooncogene Nup214 was first identified as a target for chromosomal translocation involved in leukemogenesis. Nup214 is located on the cytoplasmic face of the NPC and is implicated in anchoring the cytoplasmic filaments of the NPC and recruiting the RNA helicase Ddx19. Here, we present the crystal structure of the human Nup214 N-terminal domain at 1.65-A resolution. The structure reveals a seven-bladed beta-propeller followed by a 30-residue C-terminal extended peptide segment, which folds back onto the beta-propeller and binds to its bottom face. The beta-propeller repeats lack any recognizable sequence motif and are distinguished by extensive insertions between the canonical beta-strands. We propose a mechanism by which the C-terminal peptide extension is involved in NPC assembly.
- Laboratory of Cell Biology and Howard Hughes Medical Institute, The Rockefeller University, New York, NY 10021, USA.
Organizational Affiliation: