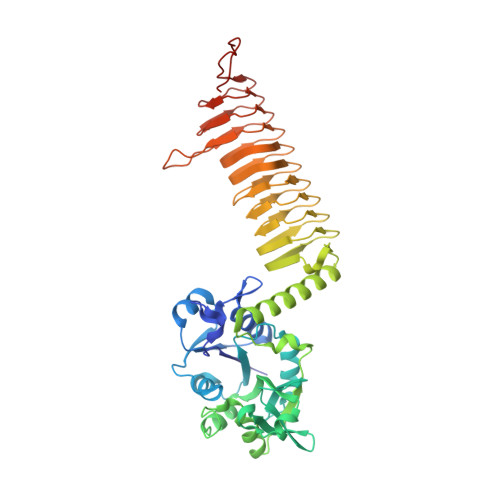
Structure of the E. coli bifunctional GlmU acetyltransferase active site with substrates and products.
Olsen, L.R., Vetting, M.W., Roderick, S.L.(2007) Protein Sci 16: 1230-1235
- PubMed: 17473010 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.072779707
- Primary Citation Related Structures:
2OI5, 2OI6, 2OI7 - PubMed Abstract:
The biosynthesis of UDP-GlcNAc in bacteria is carried out by GlmU, an essential bifunctional uridyltransferase that catalyzes the CoA-dependent acetylation of GlcN-1-PO(4) to form GlcNAc-1-PO(4) and its subsequent condensation with UTP to form pyrophosphate and UDP-GlcNAc. As a metabolite, UDP-GlcNAc is situated at a branch point leading to the biosynthesis of lipopolysaccharide and peptidoglycan. Consequently, GlmU is regarded as an important target for potential antibacterial agents. The crystal structure of the Escherichia coli GlmU acetyltransferase active site has been determined in complexes with acetyl-CoA, CoA/GlcN-1-PO(4), and desulpho-CoA/GlcNAc-1-PO(4). These structures reveal the enzyme groups responsible for binding the substrates. A superposition of these complex structures suggests that the 2-amino group of GlcN-1-PO(4) is positioned in proximity to the acetyl-CoA to facilitate direct attack on its thioester by a ternary complex mechanism.
- Department of Biochemistry, Albert Einstein College of Medicine, Bronx, New York 10461, USA.
Organizational Affiliation: