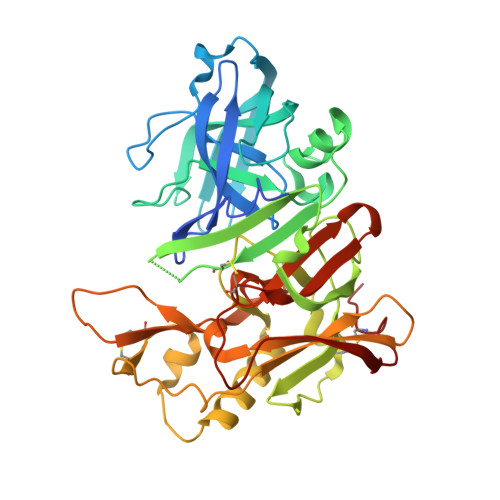
Application of fragment screening by X-ray crystallography to beta-Secretase.
Murray, C.W., Callaghan, O., Chessari, G., Cleasby, A., Congreve, M., Frederickson, M., Hartshorn, M.J., McMenamin, R., Patel, S., Wallis, N.(2007) J Med Chem 50: 1116-1123
- PubMed: 17315856 Search on PubMed
- DOI: https://doi.org/10.1021/jm0611962
- Primary Citation Related Structures:
2OF0, 2OHK, 2OHL, 2OHM, 2OHN - PubMed Abstract:
This paper describes an application of fragment screening to the aspartyl protease enzyme, beta-secretase (BACE-1), using high throughput X-ray crystallography. Three distinct chemotypes were identified by X-ray crystallography as binding to the catalytic aspartates either via an aminoheterocycle (such as 2-aminoquinoline), a piperidine, or an aliphatic hydroxyl group. The fragment hits were weak inhibitors of BACE-1 in the millimolar range but were of interest because most of them displayed relatively good ligand efficiencies. The aminoheterocycles exhibited a novel recognition motif that has not been seen before with aspartic proteases. Virtual screening around this motif identified an aminopyridine with increased potency and attractive growth points for further elaboration using structure-based drug design. The companion paper illustrates how sub-micromolar inhibitors were developed starting from this fragment.
- Astex Therapeutics Ltd., 436 Cambridge Science Park, Milton Road, Cambridge CB4 0QA, United Kingdom. c.murray@astex-therapeutics.com
Organizational Affiliation: