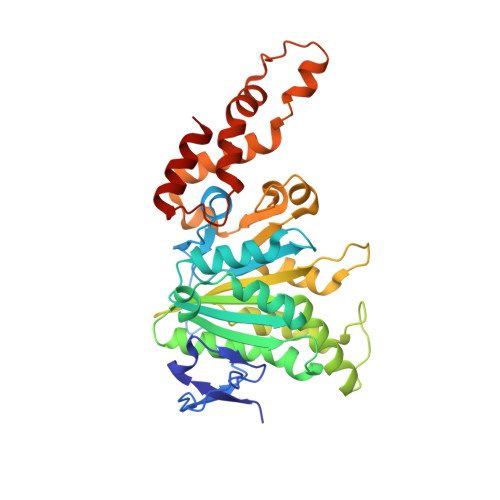
Structural analysis of a prototypical ATPase from the type III secretion system.
Zarivach, R., Vuckovic, M., Deng, W., Finlay, B.B., Strynadka, N.C.(2007) Nat Struct Mol Biol 14: 131-137
- PubMed: 17237797 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1196
- Primary Citation Related Structures:
2OBL, 2OBM - PubMed Abstract:
The type III secretion system (T3SS) ATPase is the conserved and essential inner-membrane component involved in the initial stages of selective secretion of specialized T3SS virulence effector proteins from the bacterial cytoplasm through to the infected host cell, a process crucial to subsequent pathogenicity. Here we present the 1.8-A-resolution crystal structure of the catalytic domain of the prototypical T3SS ATPase EscN from enteropathogenic Escherichia coli (EPEC). Along with in vitro and in vivo mutational analysis, our data show that the T3SS ATPases share similarity with the F1 ATPases but have important structural and sequence differences that dictate their unique secretory role. We also show that T3SS ATPase activity is dependent on EscN oligomerization and describe the molecular features and possible functional implications of a hexameric ring model.
- Department of Biochemistry and Molecular Biology and the Center for Blood Research, University of British Columbia, 2350 Health Sciences Mall, Vancouver, British Columbia V6T 1Z3, Canada.
Organizational Affiliation: