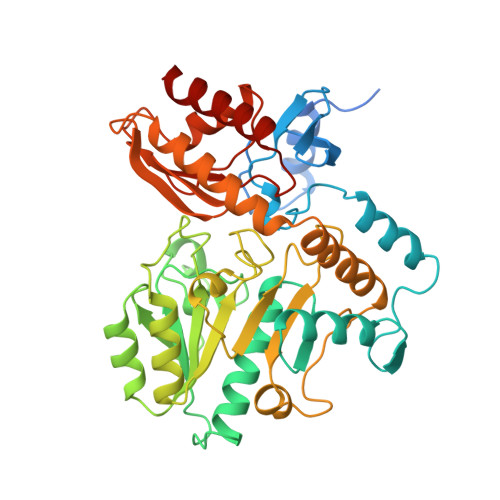
Crystal structure of human ornithine aminotransferase complexed with the highly specific and potent inhibitor 5-fluoromethylornithine.
Storici, P., Capitani, G., Muller, R., Schirmer, T., Jansonius, J.N.(1999) J Mol Biology 285: 297-309
- PubMed: 9878407 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.2289
- Primary Citation Related Structures:
2OAT - PubMed Abstract:
Ornithine aminotransferase (l-ornithine:2-oxoacid delta-aminotransferase; EC 2.6.1.13), a pyridoxal-5'-phosphate-dependent mitochondrial enzyme controls the l-ornithine level in tissues by catalyzing the transfer of the delta-amino group of l-ornithine to 2-oxoglutarate, producing l-glutamate- gamma-semialdehyde and l-glutamate. (2S, 5S)-5-Fluoromethylornithine is the only inhibitor exclusively specific for ornithine aminotransferase known to date. Both in vitro and in vivo, it blocks the enzyme by a suicide reaction leading to a covalent adduct with the cofactor. The crystal structure of the enzyme-inhibitor complex was solved at a resolution of 1.95 A. No significant conformational changes compared with the native enzyme structure were observed. The structure reveals the atomic details of the cofactor-inhibitor adduct and its interactions with the active site of the enzyme. The main residues responsible for specific binding of the inhibitor are Arg180, which forms a strong salt bridge with the alpha-carboxylate and Tyr55, which is involved in a short hydrogen bond with the alpha-amino group. The experimental observation that in the racemic mixture, (2S, 5S)-5-fluoromethylornithine is exclusively responsible for the enzyme inhibition can be explained on the basis of the active site topology. Model building studies strongly suggest that the natural substrate l-ornithine, in its external aldimine adduct with the enzyme, makes use of the same recognition site as the inhibitor. It is proposed that the neutralization of the active site Arg413 by a salt bridge with Glu235 also plays an important role in productive binding of both 5-fluoromethylornithine and l-ornithine. Arg180 and Arg413 are believed to be instrumental in recognition of l-glutamate, by binding its gamma and alpha-carboxylate groups, respectively. This requires a different side-chain conformation of Glu235. Lys292 is the only obvious candidate for catalyzing the rate-limiting proton transfer steps in the transamination reaction.
- Division of Structural Biology Biozentrum, University of Basel, Klingelbergstrasse 70, Basel, CH-4056, Switzerland.
Organizational Affiliation: