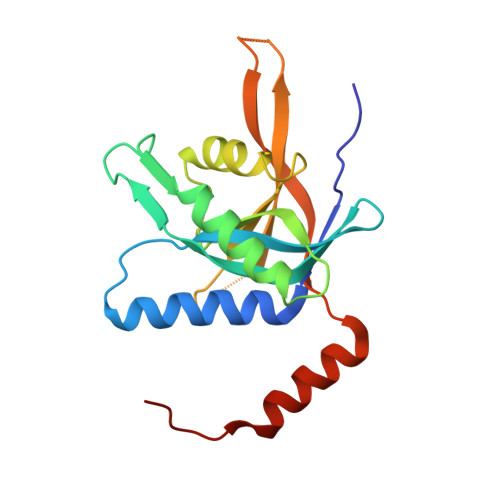
The Crystal Structure of the Human Mov34 MPN Domain Reveals a Metal-free Dimer
Sanches, M., Alves, B.S.C., Zanchin, N.I.T., Guimaraes, B.G.(2007) J Mol Biology 370: 846-855
- PubMed: 17559875 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.04.084
- Primary Citation Related Structures:
2O95, 2O96 - PubMed Abstract:
The 26S proteasome is a large protein complex involved in protein degradation. We have shown previously that the PSMD7/Mov34 subunit of the human proteasome contains a proteolytically resistant MPN domain. MPN domain family members comprise subunits of the proteasome, COP9-signalosome and translation initiation factor 3 complexes. Here, the crystal structure of two C-terminally truncated proteins, MPN 1-186 and MPN 1-177, were solved to 1.96 and 3.0 A resolution, respectively. MPN 1-186 is formed by nine beta-strands surrounded by three alpha-helices plus a fourth alpha-helix at the C terminus. This final alpha-helix emerges from the domain core and folds along with a symmetrically related subunit, typical of a domain swap. The crystallographic dimer is consistent with size-exclusion chromatography and DLS analysis showing that MPN 1-186 is a dimer in solution. MPN 1-186 shows an overall architecture highly similar to the previously reported crystal structure of the Archaeal MPN domain AfJAMM of Archaeoglobus fulgidus. However, previous structural and biophysical analyses have shown that neither MPN 1-186 nor full-length human Mov34 bind metal, in opposition to the zinc-binding AfJAMM structures. The zinc ligand residues observed in AfJAMM are conserved in the yeast Rpn11 proteasome and Csn5 COP-signalosome subunits, which is consistent with the isopeptidase activity described for these proteins. The results presented here show that, although the MPN domain of Mov34 shows a typical metalloprotease fold, it is unable to coordinate a metal ion. This finding and amino acid sequence comparisons can explain why the MPN-containing proteins Mov34/PSMD7, RPN8, Csn6, Prp8p and the translation initiation factor 3 subunits f and h do not show catalytic isopeptidase activity, allowing us to propose the hypothesis that in these proteins the MPN domain has a primarily structural function.
- Center for Structural Molecular Biology, Brazilian Synchrotron Light Laboratory (LNLS), Campinas, SP, Brazil. msanches@lnls.br
Organizational Affiliation: