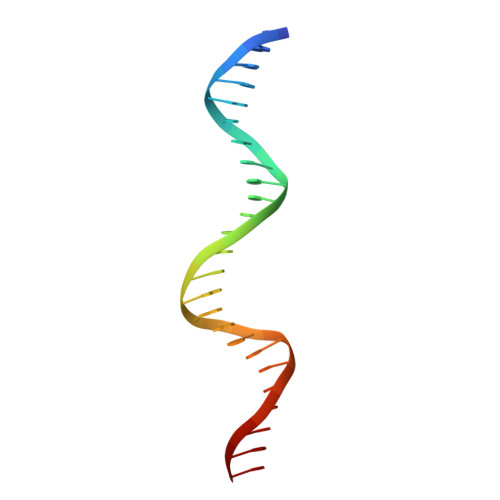
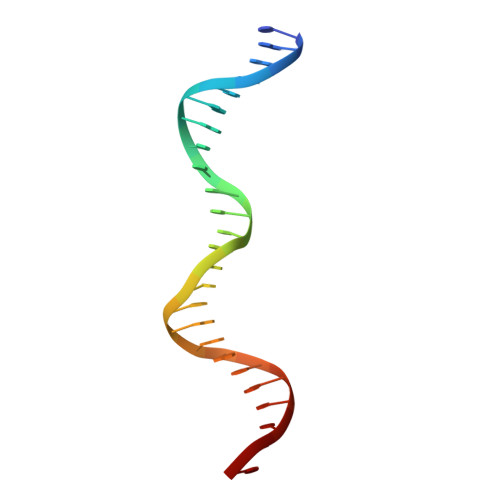
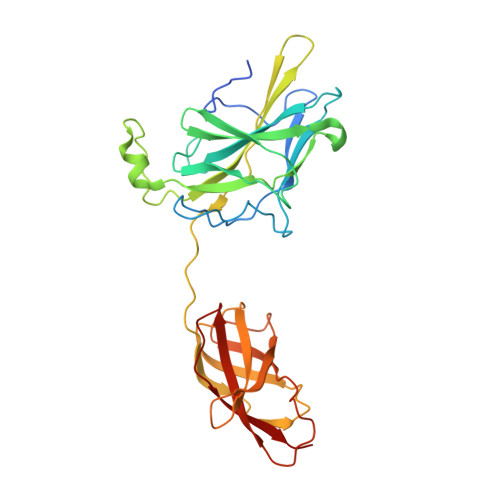
Crystal structure of NFAT bound to the HIV-1 LTR tandem kappaB enhancer element
Bates, D.L., Barthel, K.K., Wu, Y., Kalhor, R., Stroud, J.C., Giffin, M.J., Chen, L.(2008) Structure 16: 684-694
- PubMed: 18462673 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2008.01.020
- Primary Citation Related Structures:
2O93 - PubMed Abstract:
The host factor, nuclear factor of activated T-cells (NFAT), regulates the transcription and replication of HIV-1. Here, we have determined the crystal structure of the DNA binding domain of NFAT bound to the HIV-1 long terminal repeat (LTR) tandem kappaB enhancer element at 3.05 A resolution. NFAT binds as a dimer to the upstream kappaB site (Core II), but as a monomer to the 3' end of the downstream kappaB site (Core I). The DNA shows a significant bend near the 5' end of Core I, where a lysine residue from NFAT bound to the 3' end of Core II inserts into the minor groove and seems to cause DNA bases to flip out. Consistent with this structural feature, the 5' end of Core I become hypersensitive to dimethylsulfate in the in vivo footprinting upon transcriptional activation of the HIV-1 LTR. Our studies provide a basis for further investigating the functional mechanisms of NFAT in HIV-1 transcription and replication.
- Department of Chemistry and Biochemistry, University of Colorado at Boulder, Boulder, CO 80309-0215, USA.
Organizational Affiliation: