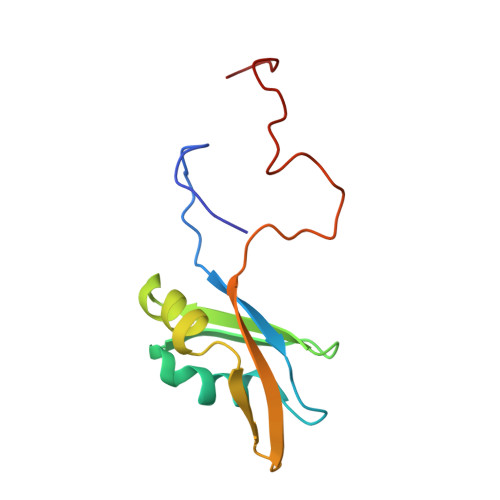
Structural and functional analysis of RNA and TAP binding to SF2/ASF.
Tintaru, A.M., Hautbergue, G.M., Hounslow, A.M., Hung, M.L., Lian, L.Y., Craven, C.J., Wilson, S.A.(2007) EMBO Rep 8: 756-762
- PubMed: 17668007 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.embor.7401031
- Primary Citation Related Structures:
2O3D - PubMed Abstract:
The serine/arginine-rich (SR) protein splicing factor 2/alternative splicing factor (SF2/ASF) has a role in splicing, stability, export and translation of messenger RNA. Here, we present the structure of the RNA recognition motif (RRM) 2 from SF2/ASF, which has an RRM fold with a considerably extended loop 5 region, containing a two-stranded beta-sheet. The loop 5 extension places the previously identified SR protein kinase 1 docking sequence largely within the RRM fold. We show that RRM2 binds to RNA in a new way, by using a tryptophan within a conserved SWQLKD motif that resides on helix alpha1, together with amino acids from strand beta2 and a histidine on loop 5. The linker connecting RRM1 and RRM2 contains arginine residues, which provide a binding site for the mRNA export factor TAP, and when TAP binds to this region it displaces RNA bound to RRM2.
- Department of Molecular Biology and Biotechnology, University of Sheffield, Firth Court, Western Bank, Sheffield S10 2TN, UK.
Organizational Affiliation: