N-(3-Cyano-4,5,6,7-tetrahydro-1-benzothien-2-yl)amides as potent, selective, inhibitors of JNK2 and JNK3.
Angell, R.M., Atkinson, F.L., Brown, M.J., Chuang, T.T., Christopher, J.A., Cichy-Knight, M., Dunn, A.K., Hightower, K.E., Malkakorpi, S., Musgrave, J.R., Neu, M., Rowland, P., Shea, R.L., Smith, J.L., Somers, D.O., Thomas, S.A., Thompson, G., Wang, R.(2007) Bioorg Med Chem Lett 17: 1296-1301
- PubMed: 17194588 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2006.12.003
- Primary Citation Related Structures:
2O0U, 2O2U - PubMed Abstract:
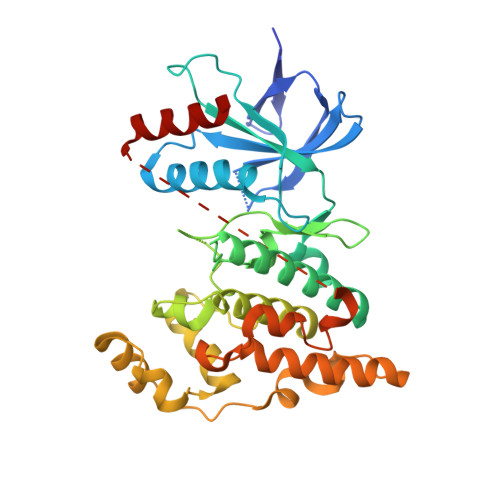
The identification and exploration of a novel, potent and selective series of N-(3-cyano-4,5,6,7-tetrahydro-1-benzothien-2-yl)amide inhibitors of JNK2 and JNK3 kinases is described. Compounds 5a and 11a were identified as potent inhibitors of JNK3 (pIC50 6.7 and 6.6, respectively), with essentially equal potency against JNK2 (pIC50 6.5). Selectivity within the mitogen-activated protein kinase (MAPK) family, against JNK1, p38alpha and ERK2, was observed for the series. X-ray crystallography of 5e and 8a in JNK3 revealed a unique binding mode, with the 3-cyano substituent forming an H-bond acceptor interaction with the hinge region of the ATP-binding site.
- GlaxoSmithKline R&D, Medicines Research Centre, Gunnels Wood Road, Stevenage, Hertfordshire SG1 2NY, UK.
Organizational Affiliation: