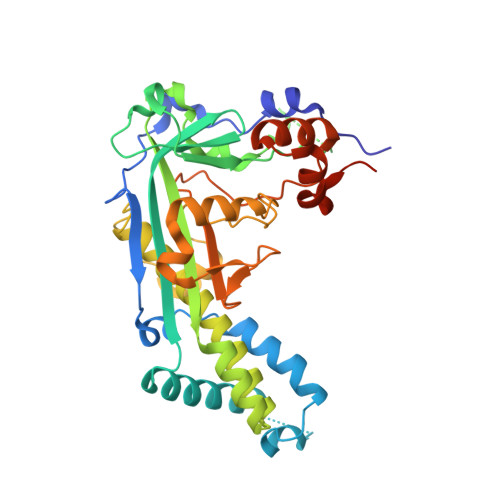
Crystal structure and solution characterization of the activation domain of human methionine synthase
Wolthers, K.R., Toogood, H.S., Jowitt, T.A., Marshall, K.R., Leys, D., Scrutton, N.S.(2007) FEBS J 274: 738-750
- PubMed: 17288554 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2006.05618.x
- Primary Citation Related Structures:
2O2K - PubMed Abstract:
Human methionine synthase (hMS) is a multidomain cobalamin-dependent enzyme that catalyses the conversion of homocysteine to methionine by methyl group transfer. We report here the 1.6 A crystal structure of the C-terminal activation domain of hMS. The structure is C-shaped with the core comprising mixed alpha and beta regions, dominated by a twisted antiparallel beta sheet with a beta-meander region. These features, including the positions of the active-site residues, are similar to the activation domain of Escherichia coli cobalamin-dependent MS (MetH). Structural and solution studies suggest a small proportion of hMS activation domain exists in a dimeric form, which contrasts with the monomeric form of the E. coli homologue. Fluorescence studies show that human activation domain interacts with the FMN-binding domain of human methionine synthase reductase (hMSR). This interaction is enhanced in the presence of S-adenosyl-methionine. Binding of the D963E/K1071N mutant activation domain to the FMN domain of MSR is weaker than with wild-type activation domain. This suggests that one or both of the residues D963 and K1071 are important in partner binding. Key differences in the sequences and structures of hMS and MetH activation domains are recognized and include a major reorientation of an extended 3(10)-containing loop in the human protein. This structural alteration might reflect differences in their respective reactivation complexes and/or potential for dimer formation. The reported structure is a component of the multidomain hMS : MSR complex, and represents an important step in understanding the impact of clinical mutations and polymorphisms in this key electron transfer complex.
- Faculty of Life Sciences, University of Manchester, UK.
Organizational Affiliation: