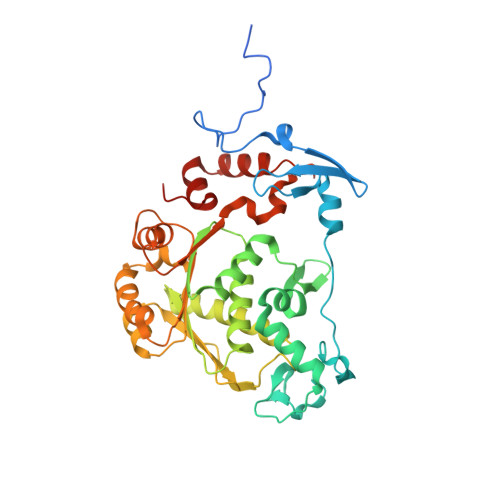
The Structure of the ATPase that Powers DNA Packaging into Bacteriophage T4 Procapsids
Sun, S., Kondabagil, K., Gentz, P.M., Rossmann, M.G., Rao, V.B.(2007) Mol Cell 25: 943-949
- PubMed: 17386269 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2007.02.013
- Primary Citation Related Structures:
2O0H, 2O0J, 2O0K - PubMed Abstract:
Packaging the viral genome into empty procapsids, an essential event in the life cycle of tailed bacteriophages and some eukaryotic viruses, is a process that shares features with chromosome assembly. Most viral procapsids possess a special vertex containing a dodecameric portal protein that is used for entry and exit of the viral genome. The portal and an ATPase are parts of the genome-packaging machine. The ATPase is required to provide energy for translocation and compaction of the negative charges on the genomic DNA. Here we report the atomic structure of the ATPase component in a phage DNA-packaging machine. The bacteriophage T4 ATPase has the greatest similarity to monomeric helicases, suggesting that the genome is translocated by an inchworm mechanism. The similarity of the packaging machines in the double-stranded DNA (dsDNA) bacteriophage T4 and dsRNA bacteriophage varphi12 is consistent with the evolution of many virions from a common ancestor.
- Department of Biological Sciences, Purdue University, 915 W. State Street, West Lafayette, IN 47907, USA.
Organizational Affiliation: