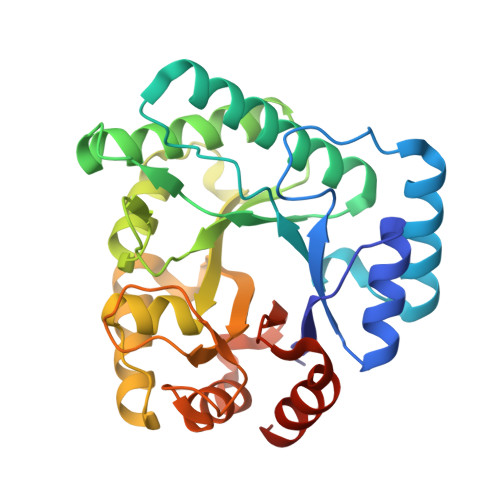
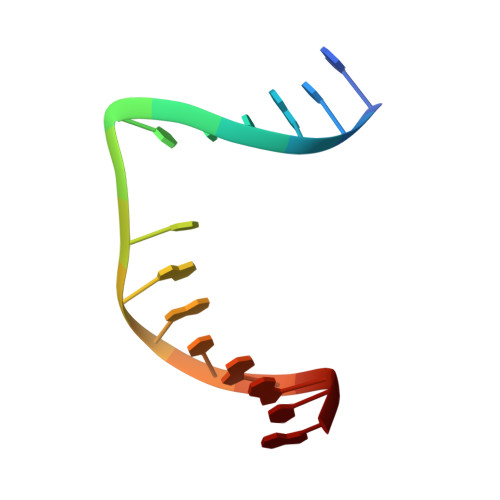
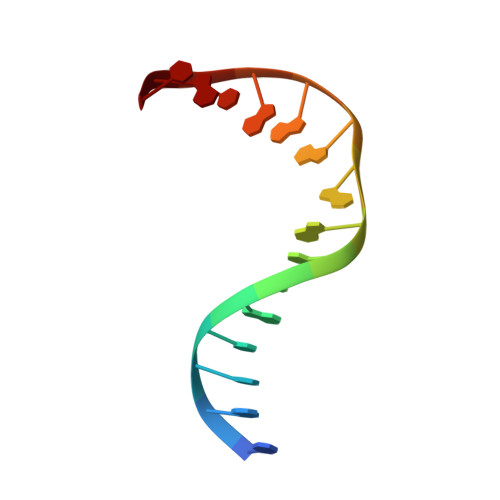
DNA apurinic-apyrimidinic site binding and excision by endonuclease IV.
Garcin, E.D., Hosfield, D.J., Desai, S.A., Haas, B.J., Bjoras, M., Cunningham, R.P., Tainer, J.A.(2008) Nat Struct Mol Biol 15: 515-522
- PubMed: 18408731 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1414
- Primary Citation Related Structures:
2NQ9, 2NQH, 2NQJ - PubMed Abstract:
Escherichia coli endonuclease IV is an archetype for an abasic or apurinic-apyrimidinic endonuclease superfamily crucial for DNA base excision repair. Here biochemical, mutational and crystallographic characterizations reveal a three-metal ion mechanism for damage binding and incision. The 1.10-A resolution DNA-free and the 2.45-A resolution DNA-substrate complex structures capture substrate stabilization by Arg37 and reveal a distorted Zn3-ligand arrangement that reverts, after catalysis, to an ideal geometry suitable to hold rather than release cleaved DNA product. The 1.45-A resolution DNA-product complex structure shows how Tyr72 caps the active site, tunes its dielectric environment and promotes catalysis by Glu261-activated hydroxide, bound to two Zn2+ ions throughout catalysis. These structural, mutagenesis and biochemical results suggest general requirements for abasic site removal in contrast to features specific to the distinct endonuclease IV alpha-beta triose phosphate isomerase (TIM) barrel and APE1 four-layer alpha-beta folds of the apurinic-apyrimidinic endonuclease families.
- Department of Molecular Biology and Skaggs Institute for Chemical Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, MB4 La Jolla, California 92037, USA.
Organizational Affiliation: