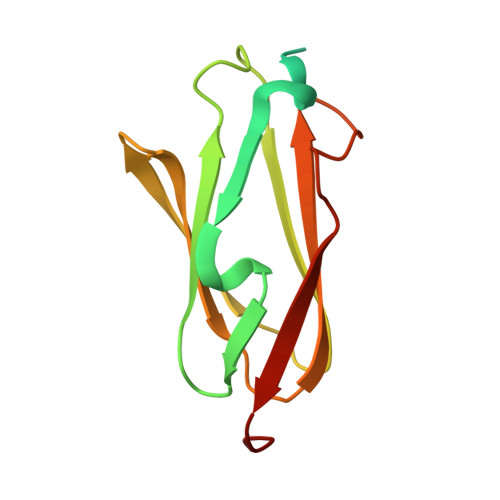
Crystal structure of human filamin C domain 23 and small angle scattering model for filamin C 23-24 dimer
Sjekloca, L., Pudas, R., Sjoeblom, B., Konarev, P., Carugo, O., Rybin, V., Kiema, T.R., Svergun, D., Ylanne, J., Djinovic-Carugo, K.(2007) J Mol Biology 368: 1011-1023
- PubMed: 17379241 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.02.018
- Primary Citation Related Structures:
2NQC - PubMed Abstract:
Filamin C is a dimeric, actin-binding protein involved in organization of cortical cytoskeleton and of the sarcomere. We performed crystallographic, small-angle X-ray scattering and analytical ultracentrifugation experiments on the constructs containing carboxy-terminal domains of the protein (domains 23-24 and 19-21). The crystal structure of domain 23 of filamin C showed that the protein adopts the expected immunoglobulin (Ig)-like fold. Small-angle X-ray scattering experiments performed on filamin C tandem Ig-like domains 23 and 24 reveal a dimer that is formed by domain 24 and that domain 23 has little interactions with itself or with domain 24, while the analytical ultracentrifugation experiments showed that the filamin C domains 19-21 form elongated monomers in diluted solutions.
- International School for Advanced Studies, 34000 Trieste, Italy.
Organizational Affiliation: