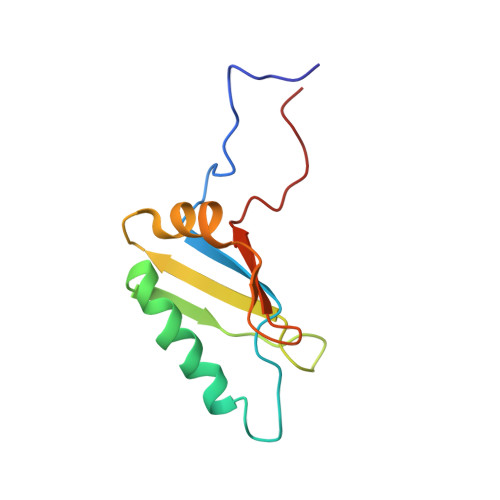
Structure of eIF3b RNA recognition motif and its interaction with eIF3j: structural insights into the recruitment of eIF3b to the 40 S ribosomal subunit.
ElAntak, L., Tzakos, A.G., Locker, N., Lukavsky, P.J.(2007) J Biological Chem 282: 8165-8174
- PubMed: 17190833 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M610860200
- Primary Citation Related Structures:
2NLW - PubMed Abstract:
Mammalian eIF3 is a 700-kDa multiprotein complex essential for initiation of protein synthesis in eukaryotic cells. It consists of 13 subunits (eIF3a to -m), among which eIF3b serves as a major scaffolding protein. Here we report the solution structure of the N-terminal RNA recognition motif of human eIF3b (eIF3b-RRM) determined by NMR spectroscopy. The structure reveals a noncanonical RRM with a negatively charged surface in the beta-sheet area contradictory with potential RNA binding activity. Instead, eIF3j, which is required for stable 40 S ribosome binding of the eIF3 complex, specifically binds to the rear alpha-helices of the eIF3b-RRM, opposite to its beta-sheet surface. Moreover, we identify that an N-terminal 69-amino acid peptide of eIF3j is sufficient for binding to eIF3b-RRM and that this interaction is essential for eIF3b-RRM recruitment to the 40 S ribosomal subunit. Our results provide the first structure of an important subdomain of a core eIF3 subunit and detailed insights into protein-protein interactions between two eIF3 subunits required for stable eIF3 recruitment to the 40 S subunit.
- Laboratory of Molecular Biology, Medical Research Council, Hills Road, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: