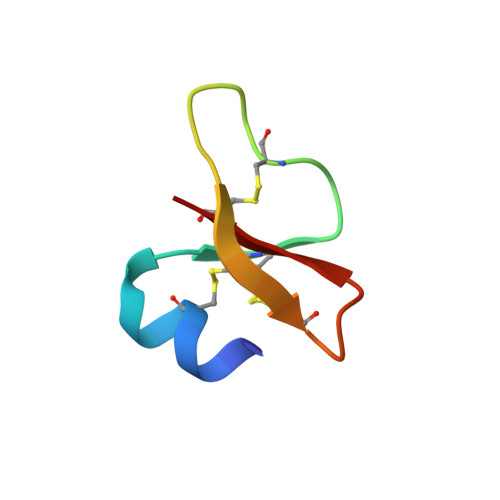
Studies of the Biological Properties of Human beta-Defensin 1.
Pazgier, M., Prahl, A., Hoover, D.M., Lubkowski, J.(2007) J Biological Chem 282: 1819-1829
- PubMed: 17071614 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M607210200
- Primary Citation Related Structures:
2NLB, 2NLC, 2NLD, 2NLE, 2NLF, 2NLG, 2NLH, 2NLP, 2NLQ, 2NLS - PubMed Abstract:
Defensins are small (30-45 amino acid residues) cationic proteins with broad antimicrobial activity against many bacteria and fungi, some enveloped viruses, and other activities such as chemoattraction of a range of different cell types to the sites of inflammation. These proteins represent attractive targets for developing novel antimicrobial agents and modulators of immune responses with therapeutic applicability. In this report, we present the results of functional and structural studies of 26 single-site mutants of human beta-defensin 1 (hBD1). All mutants were assayed for antimicrobial activity against Escherichia coli (ATCC strain 25922) and for chemotactic activity with CCR6-transfected HEK293 cells. To analyze the structural implications of mutagenesis and to verify the correctness of the disulfide connectivity, we used x-ray crystallography to conduct complete structural studies for 10 mutants in which the topology of disulfides was the same as in the native hBD1. Mutations did not induce significant changes of the tertiary structure, suggesting that the observed alterations of biological properties of the mutants were solely associated with changes in the respective side chains. We found that cationic residues located near the C terminus (Arg(29), Lys(31), Lys(33), and Lys(36)) of hBD1 define most of the anti-E. coli in vitro activity of this protein. In turn, nearly all mutations altering the CCR6-mediated chemotaxis are located at one area of the protein, defined by the N-terminal alpha-helical region (Asp(1)... Ser(8)) and a few topologically adjacent residues (Lys(22), Arg(29), and Lys(33)). These experimental results allow for the first time drafting of the CCR6-epitope for a defensin molecule.
- Macromolecular Crystallography Laboratory, NCI, National Institutes of Health, Frederick, Maryland 21702, USA.
Organizational Affiliation: