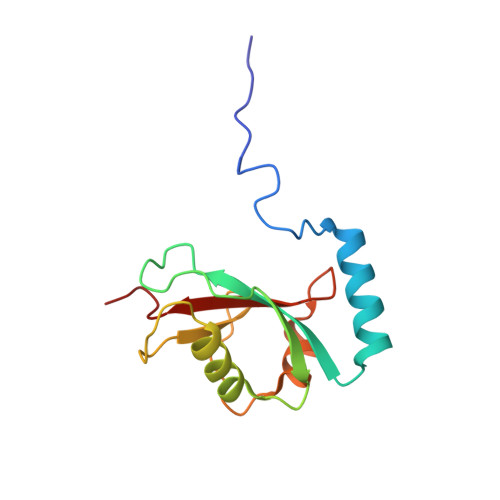
Solution structure of the autophagy-related protein LC3C reveals a polyproline II motif on a mobile tether with phosphorylation site.
Krichel, C., Mockel, C., Schillinger, O., Huesgen, P.F., Sticht, H., Strodel, B., Weiergraber, O.H., Willbold, D., Neudecker, P.(2019) Sci Rep 9: 14167-14167
- PubMed: 31578424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-48155-8
- Primary Citation Related Structures:
2NCN - PubMed Abstract:
(Macro-)autophagy is a compartmental degradation pathway conserved from yeast to mammals. The yeast protein Atg8 mediates membrane tethering/hemifusion and cargo recruitment and is essential for autophagy. The human MAP1LC3/GABARAP family proteins show high sequence identity with Atg8, but MAP1LC3C is distinguished by a conspicuous amino-terminal extension with unknown functional significance. We have determined the high-resolution three-dimensional structure and measured the backbone dynamics of MAP1LC3C by NMR spectroscopy. From Ser18 to Ala120, MAP1LC3C forms an α-helix followed by the ubiquitin-like tertiary fold with two hydrophobic binding pockets used by MAP1LC3/GABARAP proteins to recognize targets presenting LC3-interacting regions (LIRs). Unlike other MAP1LC3/GABARAP proteins, the amino-terminal region of MAP1LC3C does not form a stable helix α 1 but a "sticky arm" consisting of a polyproline II motif on a flexible linker. Ser18 at the interface between this linker and the structural core can be phosphorylated in vitro by protein kinase A, which causes additional conformational heterogeneity as monitored by NMR spectroscopy and molecular dynamics simulations, including changes in the LIR-binding interface. Based on these results we propose that the amino-terminal polyproline II motif mediates specific interactions with the microtubule cytoskeleton and that Ser18 phosphorylation modulates the interplay of MAP1LC3C with its various target proteins.
- ICS-6 (Strukturbiochemie) and JuStruct, Forschungszentrum Jülich, 52425, Jülich, Germany.
Organizational Affiliation: