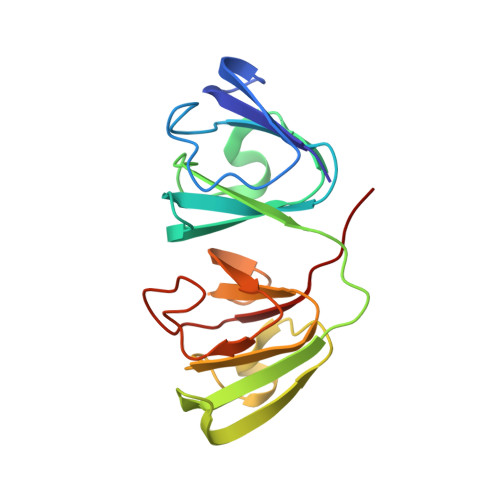
Nuclear Magnetic Resonance Structure of a Major Lens Protein, Human gamma C-Crystallin: Role of the Dipole Moment in Protein Solubility.
Dixit, K., Pande, A., Pande, J., Sarma, S.P.(2016) Biochemistry 55: 3136-3149
- PubMed: 27187112 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b00359
- Primary Citation Related Structures:
2NBR - PubMed Abstract:
A hallmark of the crystallin proteins is their exceptionally high solubility, which is vital for maintaining the high refractive index of the eye lens. Human γC-crystallin is a major γ-crystallin whose mutant forms are associated with congenital cataracts but whose three-dimensional structure is not known. An earlier study of a homology model concluded that human γC-crystallin has low intrinsic solubility, mainly because of the atypical magnitude and fluctuations of its dipole moment. On the contrary, the high-resolution tertiary structure of human γC-crystallin determined here shows unequivocally that it is a highly soluble, monomeric molecule in solution. Notable differences between the orientations and interactions of several side chains are observed upon comparison to those in the model. No evidence of the pivotal role ascribed to the effect of dipole moment on protein solubility was found. The nuclear magnetic resonance structure should facilitate a comprehensive understanding of the deleterious effects of cataract-associated mutations in human γC-crystallin.
- Molecular Biophysics Unit, Indian Institute of Science , Bangalore, Karnataka 560012, India.
Organizational Affiliation: