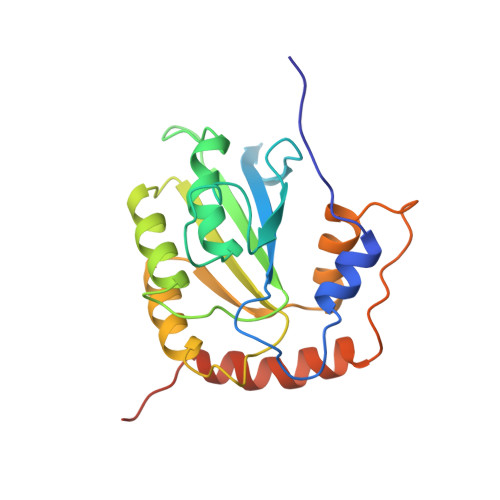
Nuclear Magnetic Resonance Structure of the APOBEC3B Catalytic Domain: Structural Basis for Substrate Binding and DNA Deaminase Activity.
Byeon, I.J., Byeon, C.H., Wu, T., Mitra, M., Singer, D., Levin, J.G., Gronenborn, A.M.(2016) Biochemistry 55: 2944-2959
- PubMed: 27163633 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b00382
- Primary Citation Related Structures:
2NBQ - PubMed Abstract:
Human APOBEC3B (A3B) is a member of the APOBEC3 (A3) family of cytidine deaminases, which function as DNA mutators and restrict viral pathogens and endogenous retrotransposons. Recently, A3B was identified as a major source of genetic heterogeneity in several human cancers. Here, we determined the solution nuclear magnetic resonance structure of the catalytically active C-terminal domain (CTD) of A3B and performed detailed analyses of its deaminase activity. The core of the structure comprises a central five-stranded β-sheet with six surrounding helices, common to all A3 proteins. The structural fold is most similar to that of A3A and A3G-CTD, with the most prominent difference being found in loop 1. The catalytic activity of A3B-CTD is ∼15-fold lower than that of A3A, although both exhibit a similar pH dependence. Interestingly, A3B-CTD with an A3A loop 1 substitution had significantly increased deaminase activity, while a single-residue change (H29R) in A3A loop 1 reduced A3A activity to the level seen with A3B-CTD. This establishes that loop 1 plays an important role in A3-catalyzed deamination by precisely positioning the deamination-targeted C into the active site. Overall, our data provide important insights into the determinants of the activities of individual A3 proteins and facilitate understanding of their biological function.
- Section on Viral Gene Regulation, Program in Genomics of Differentiation, Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health , Bethesda, Maryland 20892, United States.
Organizational Affiliation: