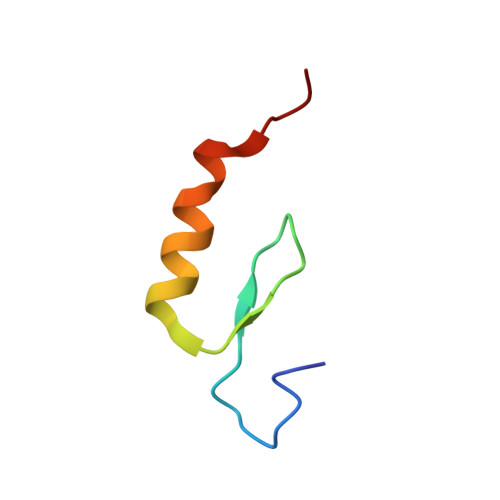
Structural basis for PHDVC5HCHNSD1-C2HRNizp1 interaction: implications for Sotos syndrome.
Berardi, A., Quilici, G., Spiliotopoulos, D., Corral-Rodriguez, M.A., Martin-Garcia, F., Degano, M., Tonon, G., Ghitti, M., Musco, G.(2016) Nucleic Acids Res 44: 3448-3463
- PubMed: 26896805 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkw103
- Primary Citation Related Structures:
2NAA, 2NAB - PubMed Abstract:
Sotos syndrome is an overgrowth syndrome caused by mutations within the functional domains ofNSD1 gene coding for NSD1, a multidomain protein regulating chromatin structure and gene expression. In particular, PHDVC5HCHNSD1 tandem domain, composed by a classical (PHDV) and an atypical (C5HCH) plant homeo-domain (PHD) finger, is target of several pathological missense-mutations. PHDVC5HCHNSD1 is also crucial for NSD1-dependent transcriptional regulation and interacts with the C2HR domain of transcriptional repressor Nizp1 (C2HRNizp1)in vitro To get molecular insights into the mechanisms dictating the patho-physiological relevance of the PHD finger tandem domain, we solved its solution structure and provided a structural rationale for the effects of seven Sotos syndrome point-mutations. To investigate PHDVC5HCHNSD1 role as structural platform for multiple interactions, we characterized its binding to histone H3 peptides and to C2HRNizp1 by ITC and NMR. We observed only very weak electrostatic interactions with histone H3 N-terminal tails, conversely we proved specific binding to C2HRNizp1 We solved C2HRNizp1 solution structure and generated a 3D model of the complex, corroborated by site-directed mutagenesis. We suggest a mechanistic scenario where NSD1 interactions with cofactors such as Nizp1 are impaired by PHDVC5HCHNSD1 pathological mutations, thus impacting on the repression of growth-promoting genes, leading to overgrowth conditions.
- Biomolecular NMR Unit, Division of Genetics and Cell Biology, IRCCS S. Raffaele Scientific Institute, Milan 20132, Italy Università degli Studi di Milano, Italy.
Organizational Affiliation: