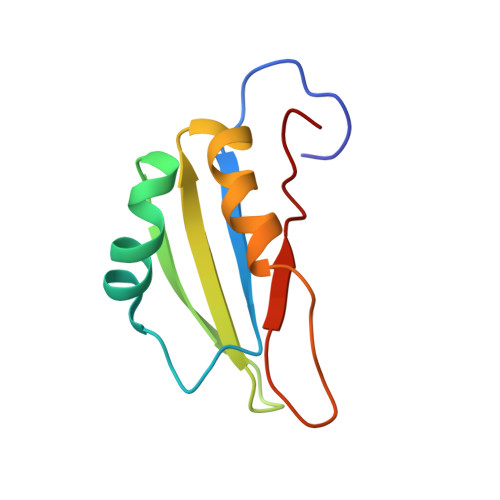
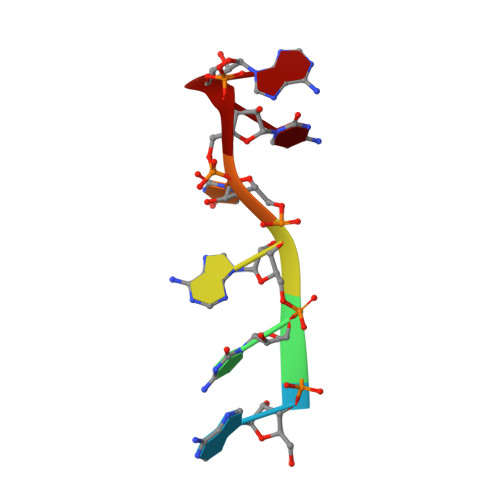
The N-terminal RNA Recognition Motif of PfSR1 Confers Semi-specificity for Pyrimidines during RNA Recognition.
Ganguly, A.K., Verma, G., Bhavesh, N.S.(2019) J Mol Biology 431: 498-510
- PubMed: 30500338 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2018.11.020
- Primary Citation Related Structures:
2N3L, 2N7C - PubMed Abstract:
Alternative splicing confers a complexity to the mRNA landscape of apicomplexans, resulting in a high proteomic diversity. The Plasmodium falciparum Ser/Arg-rich protein 1 (PfSR1) is the first protein to be confirmed as an alternative splicing factor in this class of parasitic protists [1]. A recent study [2] showed a purine bias in RNA binding among cognate RNA substrates of PfSR1. Here, we have investigated the role played by the amino-terminal RNA recognition motif (RRM1) of PfSR1 from the solution structure of its complex with ACAUCA RNA hexamer to understand how its mechanism of RNA recognition compares to human orthologs and to the C-terminal RRM. RNA binding by RRM1 is mediated through specific recognition of a cytosine base situated 5' of one or more pyrimidine bases by a conserved tyrosine residue on β 1 and a glutamate residue on the β 4 strand. Affinity is conferred through insertion of a 3' pyrimidine into a positively charged pocket. Retention of fast dynamics and ITC binding constants indicate the complex to be of moderate affinity. Using calorimetry and mapping of NMR chemical shift perturbations, we have also ascertained the purine preference of PfSR1 to be a property of the carboxy terminal pseudo-RRM (RRM2), which binds RNA non-canonically and with greater affinity compared to RRM1. Our findings show conclusive evidence of complementary RNA sequence recognition by the two RRMs, which may potentially aid PfSR1 in binding RNA with a high sequence specificity.
- Transcription Regulation group, International Centre for Genetic Engineering and Biotechnology (ICGEB), Aruna Asaf Ali Marg, New Delhi 110067, India.
Organizational Affiliation: