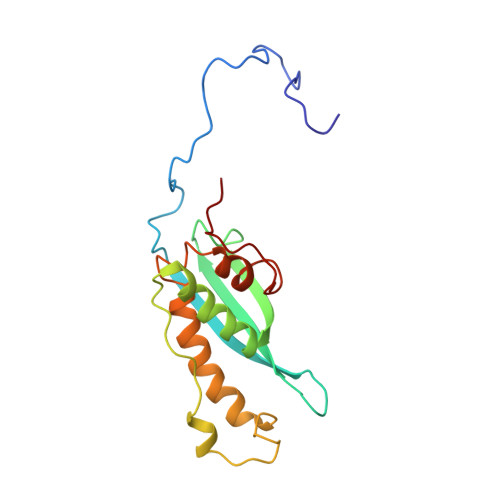
Phosphorylation of conserved phosphoinositide binding pocket regulates sorting nexin membrane targeting.
Lenoir, M., Ustunel, C., Rajesh, S., Kaur, J., Moreau, D., Gruenberg, J., Overduin, M.(2018) Nat Commun 9: 993-993
- PubMed: 29520003 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-03370-1
- Primary Citation Related Structures:
2MXC - PubMed Abstract:
Sorting nexins anchor trafficking machines to membranes by binding phospholipids. The paradigm of the superfamily is sorting nexin 3 (SNX3), which localizes to early endosomes by recognizing phosphatidylinositol 3-phosphate (PI3P) to initiate retromer-mediated segregation of cargoes to the trans-Golgi network (TGN). Here we report the solution structure of full length human SNX3, and show that PI3P recognition is accompanied by bilayer insertion of a proximal loop in its extended Phox homology (PX) domain. Phosphoinositide (PIP) binding is completely blocked by cancer-linked phosphorylation of a conserved serine beside the stereospecific PI3P pocket. This "PIP-stop" releases endosomal SNX3 to the cytosol, and reveals how protein kinases control membrane assemblies. It constitutes a widespread regulatory element found across the PX superfamily and throughout evolution including of fungi and plants. This illuminates the mechanism of a biological switch whereby structured PIP sites are phosphorylated to liberate protein machines from organelle surfaces.
- School of Cancer Sciences, College of Medical and Dental Sciences, University of Birmingham, Edgbaston, Birmingham, B15 2TT, UK.
Organizational Affiliation: