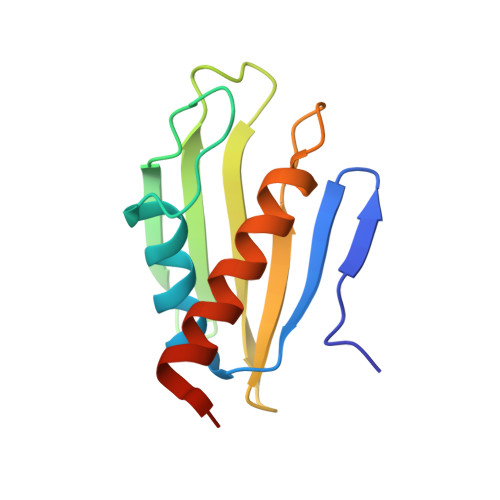
NMR Structure of Francisella tularensis Virulence Determinant Reveals Structural Homology to Bet v1 Allergen Proteins.
Zook, J., Mo, G., Sisco, N.J., Craciunescu, F.M., Hansen, D.T., Baravati, B., Cherry, B.R., Sykes, K., Wachter, R., Van Horn, W.D., Fromme, P.(2015) Structure 23: 1116-1122
- PubMed: 26004443 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.03.025
- Primary Citation Related Structures:
2MU4 - PubMed Abstract:
Tularemia is a potentially fatal bacterial infection caused by Francisella tularensis, and is endemic to North America and many parts of northern Europe and Asia. The outer membrane lipoprotein, Flpp3, has been identified as a virulence determinant as well as a potential subunit template for vaccine development. Here we present the first structure for the soluble domain of Flpp3 from the highly infectious Type A SCHU S4 strain, derived through high-resolution solution nuclear magnetic resonance (NMR) spectroscopy; the first structure of a lipoprotein from the genus Francisella. The Flpp3 structure demonstrates a globular protein with an electrostatically polarized surface containing an internal cavity-a putative binding site based on the structurally homologous Bet v1 protein family of allergens. NMR-based relaxation studies suggest loop regions that potentially modulate access to the internal cavity. The Flpp3 structure may add to the understanding of F. tularensis virulence and contribute to the development of effective vaccines.
- Department of Chemistry and Biochemistry, Arizona State University, Tempe, AZ 85287, USA; Center for Membrane Proteins in Infectious Diseases, Arizona State University, Tempe, AZ 85287, USA.
Organizational Affiliation: