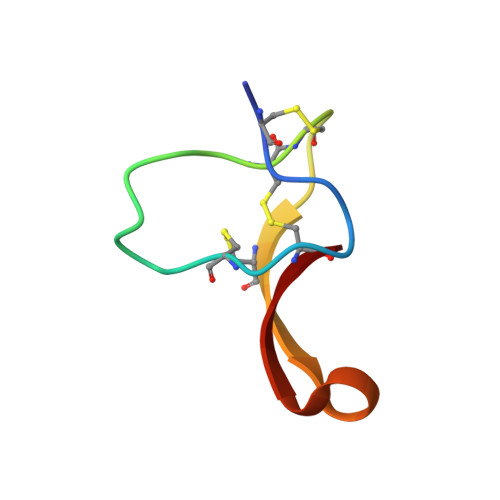
Structure of membrane-active toxin from crab spider Heriaeus melloteei suggests parallel evolution of sodium channel gating modifiers in Araneomorphae and Mygalomorphae.
Berkut, A.A., Peigneur, S., Myshkin, M.Y., Paramonov, A.S., Lyukmanova, E.N., Arseniev, A.S., Grishin, E.V., Tytgat, J., Shenkarev, Z.O., Vassilevski, A.A.(2015) J Biological Chem 290: 492-504
- PubMed: 25352595 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.595678
- Primary Citation Related Structures:
2MQU - PubMed Abstract:
We present a structural and functional study of a sodium channel activation inhibitor from crab spider venom. Hm-3 is an insecticidal peptide toxin consisting of 35 amino acid residues from the spider Heriaeus melloteei (Thomisidae). We produced Hm-3 recombinantly in Escherichia coli and determined its structure by NMR spectroscopy. Typical for spider toxins, Hm-3 was found to adopt the so-called "inhibitor cystine knot" or "knottin" fold stabilized by three disulfide bonds. Its molecule is amphiphilic with a hydrophobic ridge on the surface enriched in aromatic residues and surrounded by positive charges. Correspondingly, Hm-3 binds to both neutral and negatively charged lipid vesicles. Electrophysiological studies showed that at a concentration of 1 μm Hm-3 effectively inhibited a number of mammalian and insect sodium channels. Importantly, Hm-3 shifted the dependence of channel activation to more positive voltages. Moreover, the inhibition was voltage-dependent, and strong depolarizing prepulses attenuated Hm-3 activity. The toxin is therefore concluded to represent the first sodium channel gating modifier from an araneomorph spider and features a "membrane access" mechanism of action. Its amino acid sequence and position of the hydrophobic cluster are notably different from other known gating modifiers from spider venom, all of which are described from mygalomorph species. We hypothesize parallel evolution of inhibitor cystine knot toxins from Araneomorphae and Mygalomorphae suborders.
- From the M. M. Shemyakin and Yu. A. Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences, 117997 Moscow, Russia, Moscow Institute of Physics and Technology (State University), 117303 Moscow, Russia, and.
Organizational Affiliation: