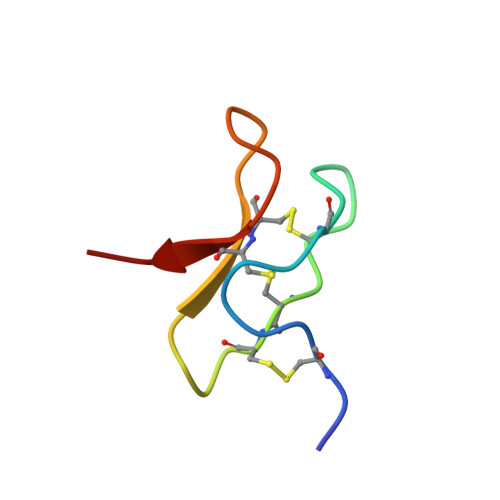
Seven novel modulators of the analgesic target NaV 1.7 uncovered using a high-throughput venom-based discovery approach.
Klint, J.K., Smith, J.J., Vetter, I., Rupasinghe, D.B., Er, S.Y., Senff, S., Herzig, V., Mobli, M., Lewis, R.J., Bosmans, F., King, G.F.(2015) Br J Pharmacol 172: 2445-2458
- PubMed: 25754331 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/bph.13081
- Primary Citation Related Structures:
2MPQ - PubMed Abstract:
Chronic pain is a serious worldwide health issue, with current analgesics having limited efficacy and dose-limiting side effects. Humans with loss-of-function mutations in the voltage-gated sodium channel NaV 1.7 (hNaV 1.7) are indifferent to pain, making hNaV 1.7 a promising target for analgesic development. Since spider venoms are replete with NaV channel modulators, we examined their potential as a source of hNaV 1.7 inhibitors. We developed a high-throughput fluorescent-based assay to screen spider venoms against hNaV 1.7 and isolate 'hit' peptides. To examine the binding site of these peptides, we constructed a panel of chimeric channels in which the S3b-S4 paddle motif from each voltage sensor domain of hNaV 1.7 was transplanted into the homotetrameric KV 2.1 channel. We screened 205 spider venoms and found that 40% contain at least one inhibitor of hNaV 1.7. By deconvoluting 'hit' venoms, we discovered seven novel members of the NaSpTx family 1. One of these peptides, Hd1a (peptide μ-TRTX-Hd1a from venom of the spider Haplopelma doriae), inhibited hNaV 1.7 with a high level of selectivity over all other subtypes, except hNaV 1.1. We showed that Hd1a is a gating modifier that inhibits hNaV 1.7 by interacting with the S3b-S4 paddle motif in channel domain II. The structure of Hd1a, determined using heteronuclear NMR, contains an inhibitor cystine knot motif that is likely to confer high levels of chemical, thermal and biological stability. Our data indicate that spider venoms are a rich natural source of hNaV 1.7 inhibitors that might be useful leads for the development of novel analgesics.
- Centre for Pain Research, Institute for Molecular Bioscience, St. Lucia, Qld, Australia.
Organizational Affiliation: