Identification, Characterization, and Three-Dimensional Structure of the Novel Circular Bacteriocin, Enterocin NKR-5-3B, from Enterococcus faecium
Himeno, K., Rosengren, K.J., Inoue, T., Perez, R.H., Colgrave, M.L., Lee, H.S., Chan, L.Y., Henriques, S.T., Fujita, K., Ishibashi, N., Zendo, T., Wilaipun, P., Nakayama, J., Leelawatcharamas, V., Jikuya, H., Craik, D.J., Sonomoto, K.(2015) Biochemistry 54: 4863-4876
- PubMed: 26174911 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.5b00196
- Primary Citation Related Structures:
2MP8 - PubMed Abstract:
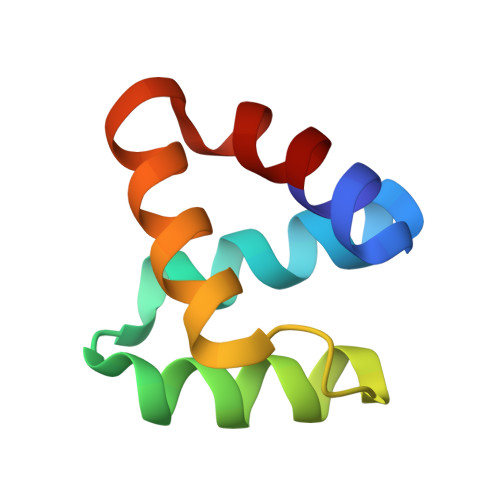
Enterocin NKR-5-3B, one of the multiple bacteriocins produced by Enterococcus faecium NKR-5-3, is a 64-amino acid novel circular bacteriocin that displays broad-spectrum antimicrobial activity. Here we report the identification, characterization, and three-dimensional nuclear magnetic resonance solution structure determination of enterocin NKR-5-3B. Enterocin NKR-5-3B is characterized by four helical segments that enclose a compact hydrophobic core, which together with its circular backbone impart high stability and structural integrity. We also report the corresponding structural gene, enkB, that encodes an 87-amino acid precursor peptide that undergoes a yet to be described enzymatic processing that involves adjacent cleavage and ligation of Leu(24) and Trp(87) to yield the mature (circular) enterocin NKR-5-3B.
- †Department of Bioscience and Biotechnology, Faculty of Agriculture, Kyushu University, 6-10-1 Hakozaki, Higashi-ku, Fukuoka 812-8581, Japan.
Organizational Affiliation: