The structure of the dimerization domain of the human polyoma, JC virus agnoprotein is an amphipathic alpha-helix
Coric, P., Saribas, S.A., Abou-Gharbia, M., Childers, W., White, M., Bouaziz, S., Safak, M.(2014) J Virol
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
(2014) J Virol
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Agnoprotein | 36 | JC polyomavirus | Mutation(s): 0 Membrane Entity: Yes | 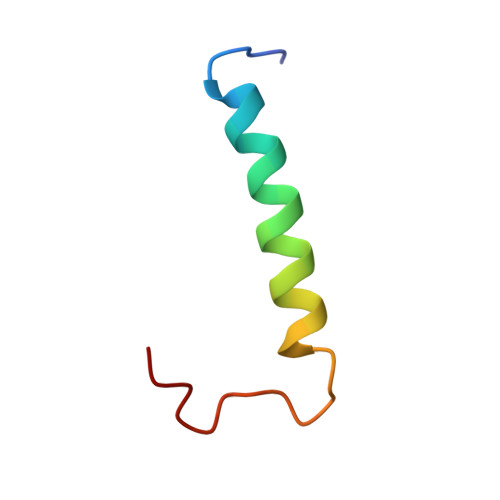 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P03086 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||