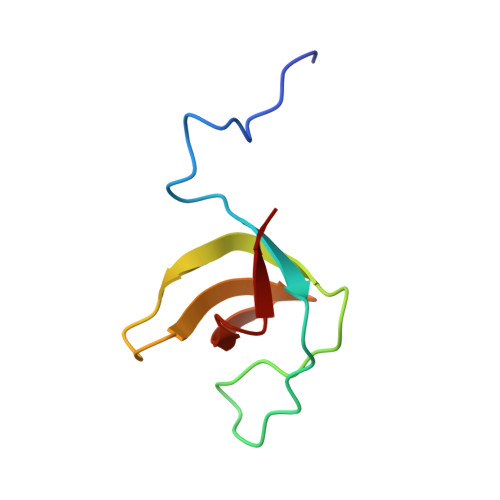
Solution NMR Structure of SH3 Domain 1 of Rho GTPase-activating Protein 10 from Homo sapiens, Northeast Structural Genomics Consortium (NESG) Target HR9129A
Xu, X., Eletsky, A., Pulavarti, S.V.S.R.K., Wang, H., O'Connell, P.T., Janjua, H., Xiao, R., Everett, J.K., Sukumaran, D., Montelione, G.T., Szyperski, T.To be published.