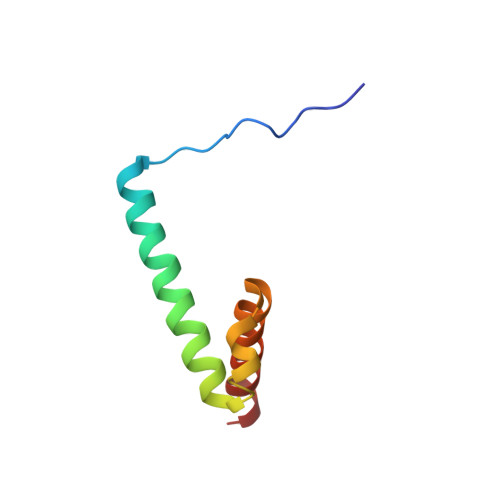
The basic helix-loop-helix region of the transcriptional repressor hairy and enhancer of split 1 is preorganized to bind DNA.
Popovic, M., Wienk, H., Coglievina, M., Boelens, R., Pongor, S., Pintar, A.(2014) Proteins 82: 537-545
- PubMed: 24403087
- DOI: https://doi.org/10.1002/prot.24507
- Primary Citation of Related Structures:
2MH3 - PubMed Abstract:
Hairy and enhancer of split 1, one of the main downstream effectors in Notch signaling, is a transcriptional repressor of the basic helix-loop-helix (bHLH) family. Using nuclear magnetic resonance methods, we have determined the structure and dynamics of a recombinant protein, H1H, which includes an N-terminal segment, b1, containing functionally important phosphorylation sites, the basic region b2, required for binding to DNA, and the HLH domain. We show that a proline residue in the sequence divides the protein in two parts, a flexible and disordered N-terminal region including b1 and a structured, mainly helical region comprising b2 and the HLH domain. Binding of H1H to a double strand DNA oligonucleotide was monitored through the chemical shift perturbation of backbone amide resonances, and showed that the interaction surface involves not only the b2 segment but also several residues in the b1 and HLH regions.
- Protein Structure and Bioinformatics Group, International Centre for Genetic Engineering and Biotechnology (ICGEB), AREA Science Park, I-34149, Trieste, Italy.
Organizational Affiliation: