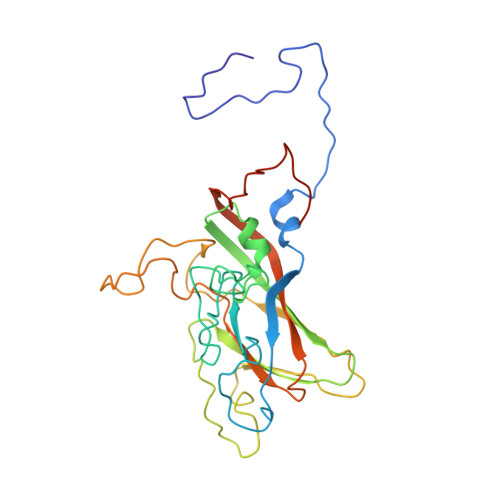
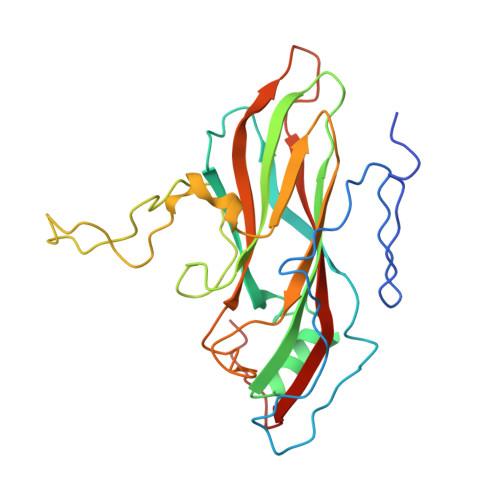
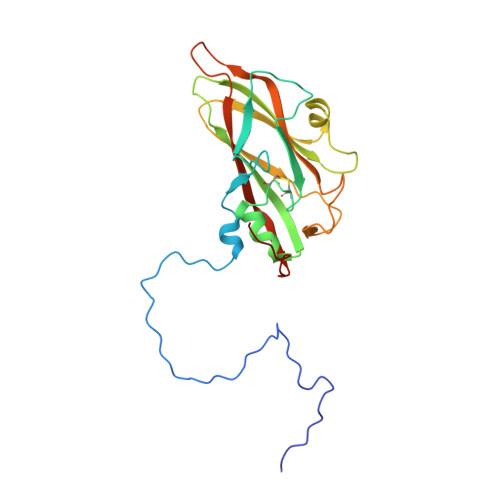
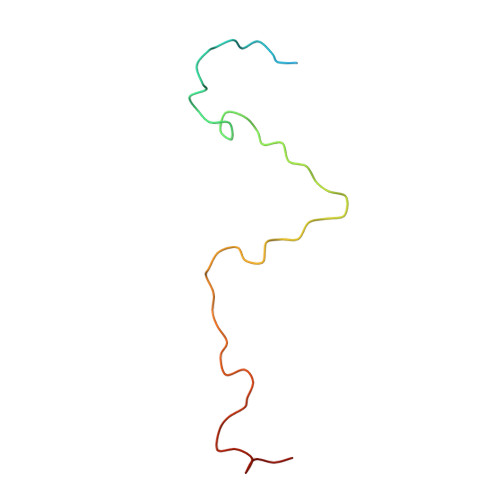
Structural refinement and analysis of Mengo virus.
Krishnaswamy, S., Rossmann, M.G.(1990) J Mol Biology 211: 803-844
- PubMed: 2156078 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(90)90077-Y
- Primary Citation Related Structures:
2MEV - PubMed Abstract:
The structure of Mengo encephalomyelitis virus was refined at 3 A resolution with a final R-factor of 0.221 and a root-mean-square deviation from idealized bond lengths of 0.019 A for 10 A to 3 A data with F greater than or equal to 3 sigma(F). The Hendrickson-Konnert refinement was restrained by the phases derived from the molecular replacement averaging procedure and constrained by the icosahedral symmetry of the virus. The virus consists of 60 protomers each having three major subunits, VP1, VP2 and VP3, along with one smaller internal protein, VP4. The three major subunits form similar eight-stranded beta-barrel structures. Alterations in the original sequence were found at position 45 in VP1 (Arg to Ala) and at position 58 in VP3 (Met to Val). The residues in loops I and II of VP1 (82 to 102), the "FMDV loop" in VP1 (205 to 213), the flexible loop of VP3 in the putative receptor attachment site (175 to 185) as well as the terminal regions 260 to 268 in VP1, 253 to 256 in VP2 and 13 to 15 in VP4 were built or modified in regions of weak density. The variation in temperature factors at the end of the refinement is over a wide range (from 2 to 80 A2), with the disordered outer and inner regions showing high mobility. Four cis proline residues, 105 in VP1, 85 and 152 in VP2 and 59 in VP3, have been identified. The disulfide bridge Cys86 to Cys88 in VP3 has been characterized. One phosphate ion and 233 water positions were included in the refinement. It is suggested that this phosphate is associated with the receptor attachment site. There are two major hydrogen-bonding networks involving solvent atoms; one involving only the subunits of a protomer, and the other connecting the protomers in a pentamer. The distribution of atom types around the icosahedral symmetry axes shows that the 5-fold channel is more hydrophobic than that along the 3-fold axis and that there are more charged residues around the 2-fold axis. The analysis of contacts between the different subunits supports the assignment of the protomeric unit. The five protomers that form the pentameric unit are held together by interactions involving the smaller VP4 protein and the amino termini of VP1 and VP3. The pentamers are associated by means of the amino-terminal region of the VP2 subunits, the beta F strand of the VP3 subunits, the C terminus of the VP4 subunits and the electrostatic helical (alpha A) interactions of VP2 subunits across the icosahedral 2-fold axes. The superposition of the corresponding subunits of Mengo virus, human rhinovirus 14 and southern bean mosaic virus has provided an improved sequence alignment. The largest structural similarity is between the VP3 subunits of Mengo virus and rhinovirus, while the least similarity is between the VP1 subunits. The various specialized insertions in the different subunits can be associated with specific functional requirements.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907.
Organizational Affiliation: