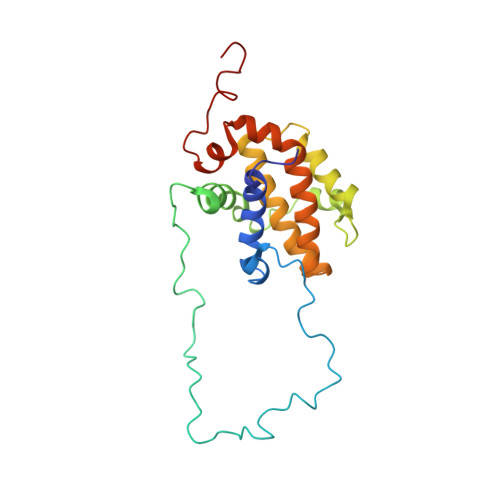
The DNA-binding domain mediates both nuclear and cytosolic functions of p53.
Follis, A.V., Llambi, F., Ou, L., Baran, K., Green, D.R., Kriwacki, R.W.(2014) Nat Struct Mol Biol 21: 535-543
- PubMed: 24814347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2829
- Primary Citation Related Structures:
2ME8, 2ME9, 2MEJ - PubMed Abstract:
Under conditions of genotoxic stress, human p53 activates the apoptotic effectors BAX or BAK to result in mitochondrial outer-membrane permeabilization and apoptosis. Antiapoptotic BCL-2 family member BCL-xL opposes this activity by sequestering cytosolic p53 via association with its DNA-binding domain, an interaction enhanced by p53 tetramerization. Here we characterized the BCL-xL-p53 complex by NMR spectroscopy and modulated it through mutagenesis to determine the relative contributions of BCL-xL's interactions with p53 or other BCL-2 family proteins to the BCL-xL-dependent inhibition of UV irradiation-induced apoptosis. Under our experimental conditions, one-third of the antiapoptotic activity of BCL-xL was mediated by p53 sequestration and the remaining two-thirds through sequestration of proapoptotic BCL-2 family members. Our studies define the contributions of cytosolic p53 to UV irradiation-induced apoptosis and provide opportunities to explore its contributions to other p53-dependent apoptotic signaling pathways.
- Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, Tennessee, USA.
Organizational Affiliation: